
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 5,719,026 – 5,719,105 |
| Length | 79 |
| Max. P | 0.870803 |

| Location | 5,719,026 – 5,719,105 |
|---|---|
| Length | 79 |
| Sequences | 10 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 82.72 |
| Shannon entropy | 0.34291 |
| G+C content | 0.46325 |
| Mean single sequence MFE | -23.79 |
| Consensus MFE | -16.90 |
| Energy contribution | -17.21 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.35 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.71 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.00 |
| SVM RNA-class probability | 0.870803 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 5719026 79 + 27905053 -AAACGAAGGCGAGCCAAUGGC-AUAAAUGGAACUGGCAUUGGCUAUGCAAAUGCAUCUAUGCA--UGCGA--GAGGGUCGCUUC---------- -......((((((.((..(.((-((..(((((....(((((.((...)).))))).)))))..)--))).)--..)).)))))).---------- ( -28.00, z-score = -2.77, R) >droSim1.chr3R 11870101 79 - 27517382 -AAACGAAGGCGAGCCAAUGGC-AUAAAUGGAACUGACAUUGGCUAUGCAAAUGCAUCUAUGCA--UGCGA--GAGGGUCGCUUC---------- -......((((((.((.(((((-...((((.......)))).)))))(((..((((....))))--)))..--..)).)))))).---------- ( -25.60, z-score = -2.44, R) >droSec1.super_0 4922052 79 - 21120651 -AAACGAAGGCGAGCCAAUGGC-AUAAAUGGAACUGACAUUGGCUAUGCAAAUGCAUCUAUGCA--UGCGA--GAGGGUCGCUUC---------- -......((((((.((.(((((-...((((.......)))).)))))(((..((((....))))--)))..--..)).)))))).---------- ( -25.60, z-score = -2.44, R) >droYak2.chr3R 9763206 79 + 28832112 -AAGCGAAGGCGAGCCAAUGGC-AUAAAUGGAACUGACAUUGGCUAUGCAAAUGCAUCUAUGCA--UGCGA--GAGGGUUGCUUC---------- -......((((((.((.(((((-...((((.......)))).)))))(((..((((....))))--)))..--..)).)))))).---------- ( -23.50, z-score = -1.24, R) >droEre2.scaffold_4770 1860139 79 - 17746568 -AAACGAAGGCGAGCCAAUGGC-AUAAAUGGAACUGACAUUGGCUAUGCAAAUGCAUCUAUGCA--UGCGA--GAGGGUUGCUUC---------- -......((((((.((.(((((-...((((.......)))).)))))(((..((((....))))--)))..--..)).)))))).---------- ( -23.50, z-score = -1.83, R) >droAna3.scaffold_13340 6280116 77 + 23697760 -UAGCGAAGGCGAGCCAAUGGC-AUAAAUGGAACUGACAUUGGCUAUGCAAAUGCAUCUAUGCA--UGCGA--GAGGGUUGCU------------ -.......(((((.((.(((((-...((((.......)))).)))))(((..((((....))))--)))..--..)).)))))------------ ( -22.80, z-score = -1.12, R) >droPer1.super_19 903460 79 - 1869541 -AAAUGAAGGCGAGCCAAUGGC-AUAAAUGGAACUGACAUUGGCUAUGCAAAUGCAUCUAUGCA--UGCGA--GAGGGUUGCCAC---------- -.......(((((.((.(((((-...((((.......)))).)))))(((..((((....))))--)))..--..)).)))))..---------- ( -26.10, z-score = -2.58, R) >dp4.chr2 25537966 79 - 30794189 -AAAUGAAGGCGAGCCAAUGGC-AUAAAUGGAACUGACAUUGGCUAUGCAAAUGCAUCUAUGCA--UGCGA--GAGGGUUGCCAC---------- -.......(((((.((.(((((-...((((.......)))).)))))(((..((((....))))--)))..--..)).)))))..---------- ( -26.10, z-score = -2.58, R) >droWil1.scaffold_181130 297003 82 - 16660200 AAACUGAAGGCGAGCAAAUUGC-AUAAAUGGAACUGGCAUUGGCCAAACGAAUGCAUUUAUGUA--UUUCAACGAGGGCUAAUCC---------- .........((((.....))))-............((.(((((((....(((((((....))))--))).......)))))))))---------- ( -16.90, z-score = -0.41, R) >droVir3.scaffold_13047 8410267 91 - 19223366 ---AAGAAGGCGUGCCAAUUGCUGUAAAUGGG-CUGACAUUGGCAAUGCUAGUGGAAAUGAGCAAGUGUCCAUAAGAGAUAAAUAAAUUAUUUGC ---.....(((((((((((.(((.......))-)....)))))).))))).(((((.((......)).)))))...((((((.....)))))).. ( -19.80, z-score = -0.36, R) >consensus _AAACGAAGGCGAGCCAAUGGC_AUAAAUGGAACUGACAUUGGCUAUGCAAAUGCAUCUAUGCA__UGCGA__GAGGGUUGCUUC__________ ........(((((.((.(((((....((((.......)))).)))))(((..((((....))))..)))......)).)))))............ (-16.90 = -17.21 + 0.31)

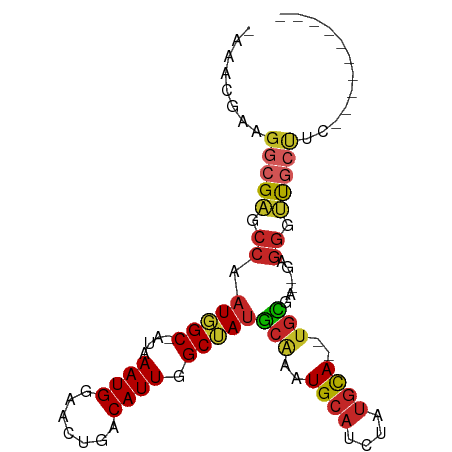
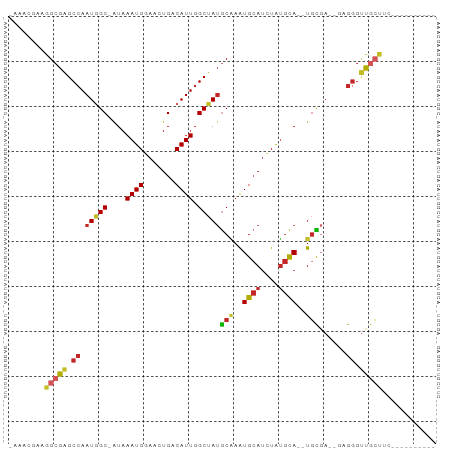
Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:01:42 2011