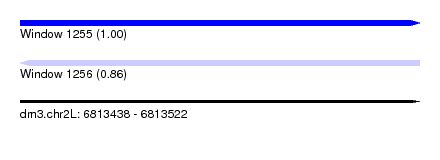

| Sequence ID | dm3.chr2L |
|---|---|
| Location | 6,813,438 – 6,813,522 |
| Length | 84 |
| Max. P | 0.997701 |

| Location | 6,813,438 – 6,813,522 |
|---|---|
| Length | 84 |
| Sequences | 11 |
| Columns | 84 |
| Reading direction | forward |
| Mean pairwise identity | 96.90 |
| Shannon entropy | 0.06413 |
| G+C content | 0.27273 |
| Mean single sequence MFE | -19.45 |
| Consensus MFE | -17.63 |
| Energy contribution | -17.72 |
| Covariance contribution | 0.09 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.89 |
| Structure conservation index | 0.91 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.16 |
| SVM RNA-class probability | 0.997701 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
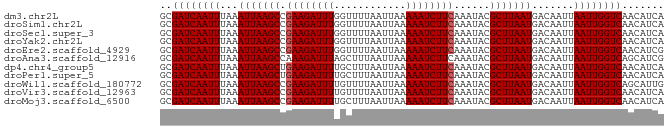
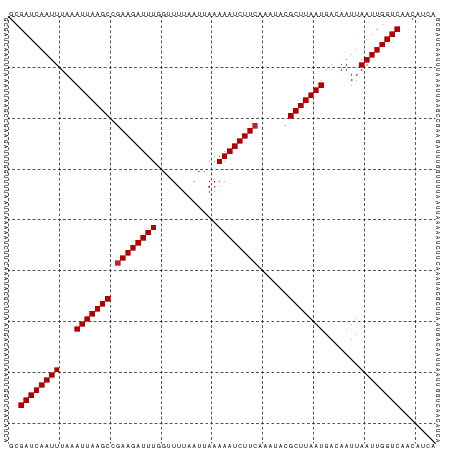
>dm3.chr2L 6813438 84 + 23011544 GCGAUCAAUUUAAAUUAAGCCGAAGAUUUGGUUUUAAUUAAAAAUCUUCAAAUACGCUUAAUGACAAUUAAUUGGUCAACAUCA ..((((((((...(((((((.((((((((............))))))))......))))))).......))))))))....... ( -18.60, z-score = -3.61, R) >droSim1.chr2L 6614351 84 + 22036055 GCGAUCAAUUUAAAUUAAGCCGAAGAUUUGGUUUUAAUUAAAAAUCUUCAAAUACGCUUAAUGACAAUUAAUUGGUCAACAUCA ..((((((((...(((((((.((((((((............))))))))......))))))).......))))))))....... ( -18.60, z-score = -3.61, R) >droSec1.super_3 2352727 84 + 7220098 GCGAUCAAUUUAAAUUAAGCCGAAGAUUUGGUUUUAAUUAAAAAUCUUCAAAUACGCUUAAUGACAAUUAAUUGGUCAACAUCA ..((((((((...(((((((.((((((((............))))))))......))))))).......))))))))....... ( -18.60, z-score = -3.61, R) >droYak2.chr2L 16230664 84 - 22324452 GCGAUCAAUUUAAAUUAAGCCGAAGAUUUGGUUUUAAUUAAAAAUCUUCAAAUACGCUUAAUGACAAUUAAUUGGUCAACAUCA ..((((((((...(((((((.((((((((............))))))))......))))))).......))))))))....... ( -18.60, z-score = -3.61, R) >droEre2.scaffold_4929 15729866 84 + 26641161 GCGAUCAAUUUAAAUUAAGCCGAAGAUUUGGUUUUAAUUAAAAAUCUUCAAAUACGCUUAAUGACAAUUAAUUGGUCAACAUCG ..((((((((...(((((((.((((((((............))))))))......))))))).......))))))))....... ( -18.60, z-score = -3.47, R) >droAna3.scaffold_12916 8398921 84 + 16180835 GCGAUCAAUUUAAAUUAAGCCAAAGAUUUAGCUUUAAUUAAAAAUCUUCAAAUACGCUUAAUGACAAUUAAUUGGUCAGCAUCG ((((((((((...(((((((..(((((((............))))))).......))))))).......)))))))).)).... ( -16.10, z-score = -2.50, R) >dp4.chr4_group5 297798 84 + 2436548 GCGAUCAAUUUAAAUUAAGCUGAAGAUUUUGCUUUAAUUAAAAAUCUUCAAAUACGCUUAAUGACAAUUAAUUGGUCAACAUCA ..((((((((...(((((((((((((((((..........)))))))))).....))))))).......))))))))....... ( -22.00, z-score = -4.95, R) >droPer1.super_5 4795223 84 + 6813705 GCGAUCAAUUUAAAUUAAGCUGAAGAUUUUGCUUUAAUUAAAAAUCUUCAAAUACGCUUAAUGACAAUUAAUUGGUCAACAUCA ..((((((((...(((((((((((((((((..........)))))))))).....))))))).......))))))))....... ( -22.00, z-score = -4.95, R) >droWil1.scaffold_180772 6952467 84 + 8906247 GCGAUCAAUUUAAAUUAAGCCGAAGAUUUUGUUUUAAUUAAAAAUCUUCAAAUACGCUUAAUGACAAUUAAUUGGUCAGCAUUG ((((((((((...(((((((.(((((((((..........)))))))))......))))))).......)))))))).)).... ( -21.40, z-score = -4.11, R) >droVir3.scaffold_12963 5328859 84 - 20206255 GCGAUCAAUUUAAAUUAAGCCGAAGAUUUUGUUUUAAUUAAAAAUCUUCAAAUACGCUUAAUGACAAUUAAUUGGUCAACAUCA ..((((((((...(((((((.(((((((((..........)))))))))......))))))).......))))))))....... ( -19.70, z-score = -4.26, R) >droMoj3.scaffold_6500 14520012 84 - 32352404 GCGAUCAAUUUAAAUUAAGCCGAAGAUUUUGCUUUAAUUAAAAAUCUUCAAAUACGCUUAAUGACAAUUAAUUGGUCAACAUCA ..((((((((...(((((((.(((((((((..........)))))))))......))))))).......))))))))....... ( -19.70, z-score = -4.11, R) >consensus GCGAUCAAUUUAAAUUAAGCCGAAGAUUUGGUUUUAAUUAAAAAUCUUCAAAUACGCUUAAUGACAAUUAAUUGGUCAACAUCA ..((((((((...(((((((.((((((((............))))))))......))))))).......))))))))....... (-17.63 = -17.72 + 0.09)
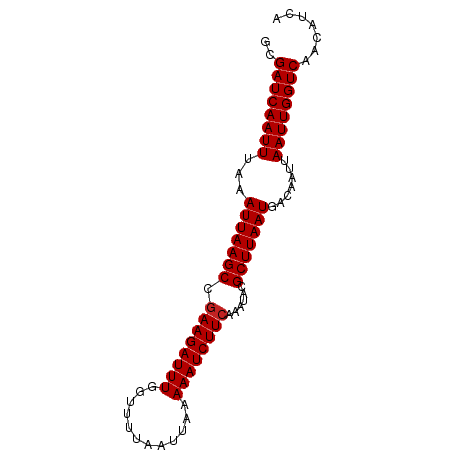

| Location | 6,813,438 – 6,813,522 |
|---|---|
| Length | 84 |
| Sequences | 11 |
| Columns | 84 |
| Reading direction | reverse |
| Mean pairwise identity | 96.90 |
| Shannon entropy | 0.06413 |
| G+C content | 0.27273 |
| Mean single sequence MFE | -15.15 |
| Consensus MFE | -14.86 |
| Energy contribution | -14.63 |
| Covariance contribution | -0.23 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.42 |
| Structure conservation index | 0.98 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.97 |
| SVM RNA-class probability | 0.863847 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
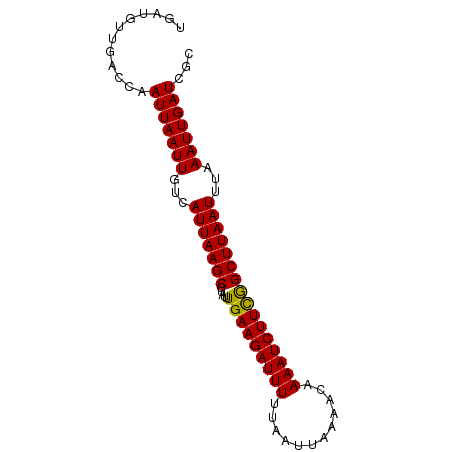
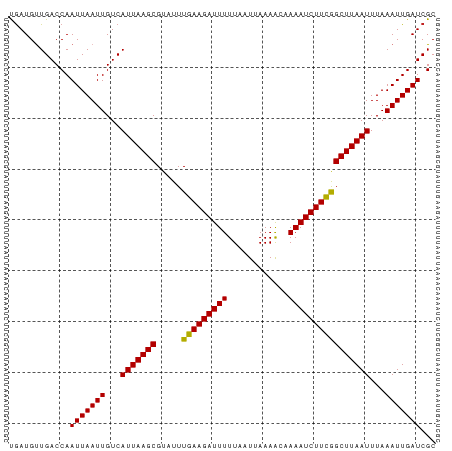
>dm3.chr2L 6813438 84 - 23011544 UGAUGUUGACCAAUUAAUUGUCAUUAAGCGUAUUUGAAGAUUUUUAAUUAAAACCAAAUCUUCGGCUUAAUUUAAAUUGAUCGC ............(((((((...(((((((.....(((((((((............))))))))))))))))...)))))))... ( -14.60, z-score = -1.39, R) >droSim1.chr2L 6614351 84 - 22036055 UGAUGUUGACCAAUUAAUUGUCAUUAAGCGUAUUUGAAGAUUUUUAAUUAAAACCAAAUCUUCGGCUUAAUUUAAAUUGAUCGC ............(((((((...(((((((.....(((((((((............))))))))))))))))...)))))))... ( -14.60, z-score = -1.39, R) >droSec1.super_3 2352727 84 - 7220098 UGAUGUUGACCAAUUAAUUGUCAUUAAGCGUAUUUGAAGAUUUUUAAUUAAAACCAAAUCUUCGGCUUAAUUUAAAUUGAUCGC ............(((((((...(((((((.....(((((((((............))))))))))))))))...)))))))... ( -14.60, z-score = -1.39, R) >droYak2.chr2L 16230664 84 + 22324452 UGAUGUUGACCAAUUAAUUGUCAUUAAGCGUAUUUGAAGAUUUUUAAUUAAAACCAAAUCUUCGGCUUAAUUUAAAUUGAUCGC ............(((((((...(((((((.....(((((((((............))))))))))))))))...)))))))... ( -14.60, z-score = -1.39, R) >droEre2.scaffold_4929 15729866 84 - 26641161 CGAUGUUGACCAAUUAAUUGUCAUUAAGCGUAUUUGAAGAUUUUUAAUUAAAACCAAAUCUUCGGCUUAAUUUAAAUUGAUCGC ............(((((((...(((((((.....(((((((((............))))))))))))))))...)))))))... ( -14.60, z-score = -1.27, R) >droAna3.scaffold_12916 8398921 84 - 16180835 CGAUGCUGACCAAUUAAUUGUCAUUAAGCGUAUUUGAAGAUUUUUAAUUAAAGCUAAAUCUUUGGCUUAAUUUAAAUUGAUCGC ((((.......((((((..(((.(((((....))))).)))..)))))).(((((((....)))))))...........)))). ( -13.70, z-score = -0.38, R) >dp4.chr4_group5 297798 84 - 2436548 UGAUGUUGACCAAUUAAUUGUCAUUAAGCGUAUUUGAAGAUUUUUAAUUAAAGCAAAAUCUUCAGCUUAAUUUAAAUUGAUCGC ............(((((((...(((((((.....((((((((((..........)))))))))))))))))...)))))))... ( -16.40, z-score = -1.77, R) >droPer1.super_5 4795223 84 - 6813705 UGAUGUUGACCAAUUAAUUGUCAUUAAGCGUAUUUGAAGAUUUUUAAUUAAAGCAAAAUCUUCAGCUUAAUUUAAAUUGAUCGC ............(((((((...(((((((.....((((((((((..........)))))))))))))))))...)))))))... ( -16.40, z-score = -1.77, R) >droWil1.scaffold_180772 6952467 84 - 8906247 CAAUGCUGACCAAUUAAUUGUCAUUAAGCGUAUUUGAAGAUUUUUAAUUAAAACAAAAUCUUCGGCUUAAUUUAAAUUGAUCGC ............(((((((...(((((((.....((((((((((..........)))))))))))))))))...)))))))... ( -15.70, z-score = -1.82, R) >droVir3.scaffold_12963 5328859 84 + 20206255 UGAUGUUGACCAAUUAAUUGUCAUUAAGCGUAUUUGAAGAUUUUUAAUUAAAACAAAAUCUUCGGCUUAAUUUAAAUUGAUCGC ............(((((((...(((((((.....((((((((((..........)))))))))))))))))...)))))))... ( -15.70, z-score = -1.74, R) >droMoj3.scaffold_6500 14520012 84 + 32352404 UGAUGUUGACCAAUUAAUUGUCAUUAAGCGUAUUUGAAGAUUUUUAAUUAAAGCAAAAUCUUCGGCUUAAUUUAAAUUGAUCGC ............(((((((...(((((((.....((((((((((..........)))))))))))))))))...)))))))... ( -15.70, z-score = -1.29, R) >consensus UGAUGUUGACCAAUUAAUUGUCAUUAAGCGUAUUUGAAGAUUUUUAAUUAAAACAAAAUCUUCGGCUUAAUUUAAAUUGAUCGC ............(((((((...(((((((.....(((((((((............))))))))))))))))...)))))))... (-14.86 = -14.63 + -0.23)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:22:10 2011