
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 4,996,889 – 4,996,988 |
| Length | 99 |
| Max. P | 0.740971 |

| Location | 4,996,889 – 4,996,988 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 92.36 |
| Shannon entropy | 0.13151 |
| G+C content | 0.49471 |
| Mean single sequence MFE | -17.13 |
| Consensus MFE | -14.38 |
| Energy contribution | -14.82 |
| Covariance contribution | 0.44 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.85 |
| Structure conservation index | 0.84 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.740971 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
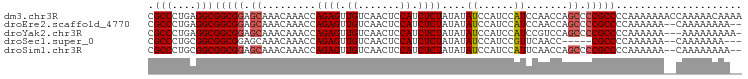
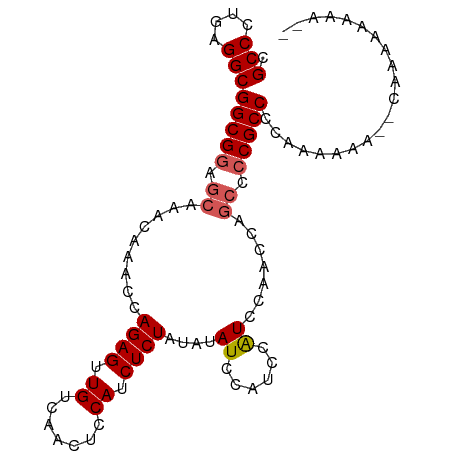
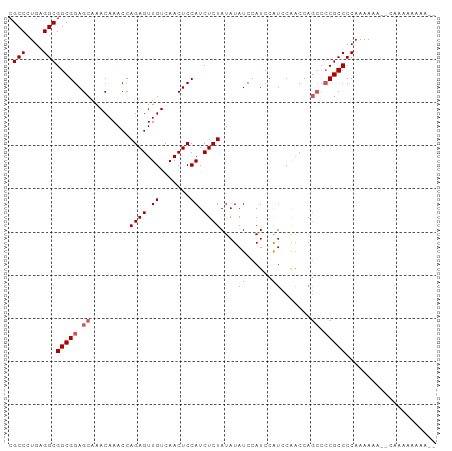
>dm3.chr3R 4996889 99 + 27905053 CGCCCUGAGGCGGCGGAGCAAACAAACCAGAGUUGUCAACUCCAUCUCUAUAUAUCCAUCCAUCCAACCAGCCCCGCCCCAAAAAAACCAAAAACAAAA .(((....)))(((((.((.........((((.((.......)).))))((......))...........)).)))))..................... ( -17.10, z-score = -2.00, R) >droEre2.scaffold_4770 1141589 95 - 17746568 CGCCCUGAGGCGGCGGAGCAAACAAACCAGAGUUGUCAACUCCAUCUCUAUAUAUCCAUCCAUCCAACCAGCCCCGCCCCAAAAAA--CAAAAAAAA-- .(((....)))(((((.((.........((((.((.......)).))))((......))...........)).)))))........--.........-- ( -17.10, z-score = -1.97, R) >droYak2.chr3R 9044823 95 + 28832112 CGCCCUGAGGCGGCGGAGCAAACAAACCAGAGUUGUCAACUCCAUCUCUAUAUAUCCAUCCAUCCGUCCAGCCCCGCCCCAAAAAA---AAAAAAAAA- .(((....)))(((((.((.........((((.((.......)).))))...........(....)....)).)))))........---.........- ( -18.10, z-score = -2.17, R) >droSec1.super_0 4235475 89 - 21120651 CGCCCUGCGGCGGCGGAGCAAACAAACCAGAGUUGUCAACUCCAUCUCUAUAUAUCCAUCCGUUCAACC-----CGCCCCAAAAAA--CAAAAAAA--- .....((.(((((((((...........((((.((.......)).)))).........))))......)-----)))).)).....--........--- ( -16.35, z-score = -1.46, R) >droSim1.chr3R 11201938 95 - 27517382 CGCCCUGCGGCGGCGGAGCAAACAAACCAGAGUUGUCAACUCCAUCUCUAUAUAUCCAUCCAUUCAACCAGCCCCGCCCCAAAAAA--CAAAAAAAA-- .(((....)))(((((.((.........((((.((.......)).))))((......))...........)).)))))........--.........-- ( -17.00, z-score = -1.67, R) >consensus CGCCCUGAGGCGGCGGAGCAAACAAACCAGAGUUGUCAACUCCAUCUCUAUAUAUCCAUCCAUCCAACCAGCCCCGCCCCAAAAAA__CAAAAAAAA__ .(((....)))(((((.((.........((((.((.......)).))))....((......)).......)).)))))..................... (-14.38 = -14.82 + 0.44)
| Location | 4,996,889 – 4,996,988 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 92.36 |
| Shannon entropy | 0.13151 |
| G+C content | 0.49471 |
| Mean single sequence MFE | -29.68 |
| Consensus MFE | -25.56 |
| Energy contribution | -26.36 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.86 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.30 |
| SVM RNA-class probability | 0.636476 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
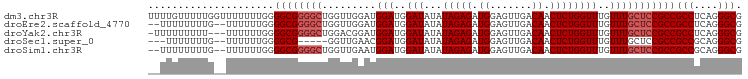
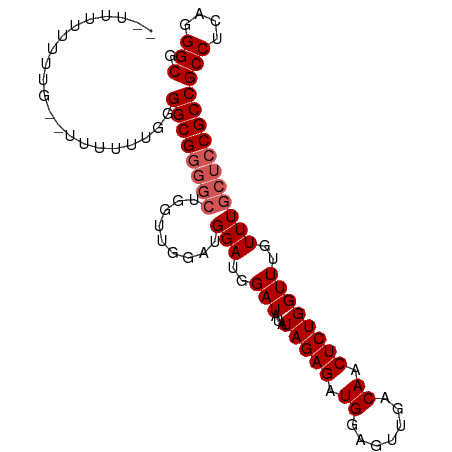
>dm3.chr3R 4996889 99 - 27905053 UUUUGUUUUUGGUUUUUUUGGGGCGGGGCUGGUUGGAUGGAUGGAUAUAUAGAGAUGGAGUUGACAACUCUGGUUUGUUUGCUCCGCCGCCUCAGGGCG .....................((((((((.........(((..(((...(((((.((.......)).))))))))..)))))))))))(((....))). ( -28.60, z-score = -0.75, R) >droEre2.scaffold_4770 1141589 95 + 17746568 --UUUUUUUUG--UUUUUUGGGGCGGGGCUGGUUGGAUGGAUGGAUAUAUAGAGAUGGAGUUGACAACUCUGGUUUGUUUGCUCCGCCGCCUCAGGGCG --........(--((((..((((((((((.........(((..(((...(((((.((.......)).))))))))..))))))))))).))..))))). ( -29.40, z-score = -1.38, R) >droYak2.chr3R 9044823 95 - 28832112 -UUUUUUUUU---UUUUUUGGGGCGGGGCUGGACGGAUGGAUGGAUAUAUAGAGAUGGAGUUGACAACUCUGGUUUGUUUGCUCCGCCGCCUCAGGGCG -.........---........((((((((.((((((((...........(((((.((.......)).)))))))))))))))))))))(((....))). ( -31.30, z-score = -2.18, R) >droSec1.super_0 4235475 89 + 21120651 ---UUUUUUUG--UUUUUUGGGGCG-----GGUUGAACGGAUGGAUAUAUAGAGAUGGAGUUGACAACUCUGGUUUGUUUGCUCCGCCGCCGCAGGGCG ---.......(--((((.(((((((-----((..((((((((.......(((((.((.......)).)))))))))))))..)))))).))).))))). ( -28.81, z-score = -1.98, R) >droSim1.chr3R 11201938 95 + 27517382 --UUUUUUUUG--UUUUUUGGGGCGGGGCUGGUUGAAUGGAUGGAUAUAUAGAGAUGGAGUUGACAACUCUGGUUUGUUUGCUCCGCCGCCGCAGGGCG --........(--((((.(((((((((((.........(((..(((...(((((.((.......)).))))))))..))))))))))).))).))))). ( -30.30, z-score = -1.73, R) >consensus __UUUUUUUUG__UUUUUUGGGGCGGGGCUGGUUGGAUGGAUGGAUAUAUAGAGAUGGAGUUGACAACUCUGGUUUGUUUGCUCCGCCGCCUCAGGGCG .....................((((((((.........(((..(((...(((((.((.......)).))))))))..)))))))))))(((....))). (-25.56 = -26.36 + 0.80)

Generated by rnazCluster.pl (part of RNAz 1.0) on Wed Apr 20 00:00:11 2011