
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 6,784,017 – 6,784,116 |
| Length | 99 |
| Max. P | 0.763936 |

| Location | 6,784,017 – 6,784,116 |
|---|---|
| Length | 99 |
| Sequences | 8 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 73.58 |
| Shannon entropy | 0.48360 |
| G+C content | 0.37330 |
| Mean single sequence MFE | -28.16 |
| Consensus MFE | -11.01 |
| Energy contribution | -11.39 |
| Covariance contribution | 0.38 |
| Combinations/Pair | 1.28 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.39 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.62 |
| SVM RNA-class probability | 0.763936 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
>dm3.chr2L 6784017 99 + 23011544 ----------UAAAUUAGCUUAAGCAAAUUAGGCUAGUUUCCUGCGC--UUUAAUUGA-----AUUGUGUAUUUUAU-CGCAGACUU-GUCGCUAUCGUGACAAUUAGUUAGCAGCAU ----------.((((((((((((.....)))))))))))).....((--(((((((((-----.(((((........-)))))..((-(((((....))))))))))))))).))).. ( -29.50, z-score = -3.03, R) >droPer1.super_5 5434554 115 - 6813705 UGCAUUGGGUUCGACAUUUGUACGCAAAUUAGAGCUACUUCCUGUGG--CUUAAUUGAGAUGAAAUGUUUGCUUUGGGCCCUGACUUUGUCGCUAUCGUGACAAUUAGUUAGCGACU- .((...((((((((.(((((....)))))..(((((((.....))))--))).....................))))))))((((((((((((....)))))))..)))))))....- ( -34.00, z-score = -2.22, R) >dp4.chr4_group1 3891254 115 + 5278887 UGCAUUGGGUUCGACAUUUGUACGCAAAUUAGAGCUACUUCCUGUGG--CUUAAUUGAGAUGAAAUGUUUGCUUUGGGCGCUGACUUUGUCGCUAUCGUGACAAUUAGUUAGCGACU- .((...((((..(((((((...(...((((.(((((((.....))))--)))))))..)...))))))).))))...))((((((((((((((....)))))))..)))))))....- ( -37.30, z-score = -2.95, R) >droAna3.scaffold_12916 8362675 101 + 16180835 -------------GAGAAUCAAAGUAAAUGAAUGUCAGUCUAGGUAAAACUUAAUUAAGU---UGUGUGCAUUUUAU-CACCGACUUUCCCGCUAUCGUGACAAUUAGUUAGCAGAAC -------------..........((.(((...(((((...((((.....))))...((((---((.(((........-)))))))))...........)))))....))).))..... ( -13.80, z-score = 0.88, R) >droEre2.scaffold_4929 15700596 99 + 26641161 ----------UAAAUUAGCUUAAGCAAAUUAGGCUCGUUUCCUGCGC--UUUAAUUGA-----AUUGUGUAUUUUAU-CGCAGACUU-GUCGCUAUCGUGACAAUUAGUUAGCAGCAU ----------.((((.(((((((.....))))))).)))).....((--(((((((((-----.(((((........-)))))..((-(((((....))))))))))))))).))).. ( -25.90, z-score = -1.82, R) >droYak2.chr2L 16199619 99 - 22324452 ----------UAAAUUAGCUUAAGCAAAUUAGGCUCGUUUCCUGCGC--UUUAAUUGA-----AUUGUGUAUUUUAU-CGCAGACUU-GUCGCUAUCGUGACAAUUAGUUAGCAGCAU ----------.((((.(((((((.....))))))).)))).....((--(((((((((-----.(((((........-)))))..((-(((((....))))))))))))))).))).. ( -25.90, z-score = -1.82, R) >droSec1.super_3 2322810 99 + 7220098 ----------UAAAUUAGCUUAAGCAAAUUAGGCUAGUUUCCUGCGC--UUUAAUUAA-----AUUGUGUAUUUUAU-CGCAGACUU-GUCGCUAUCGUGACAAUUAGUUAGCAGCAU ----------.((((((((((((.....)))))))))))).....((--(((((((((-----.(((((........-)))))..((-(((((....))))))))))))))).))).. ( -29.40, z-score = -3.35, R) >droSim1.chr2L 6585261 99 + 22036055 ----------UAAAUUAGCUUAAGCAAAUUAGGCUAGUUUCCUGCGC--UUUAAUUGA-----AUUGUGUAUUUUAU-CGCAGACUU-GUCGCUAUCGUGACAAUUAGUUAGCAGCAU ----------.((((((((((((.....)))))))))))).....((--(((((((((-----.(((((........-)))))..((-(((((....))))))))))))))).))).. ( -29.50, z-score = -3.03, R) >consensus __________UAAAUUAGCUUAAGCAAAUUAGGCUAGUUUCCUGCGC__UUUAAUUGA_____AUUGUGUAUUUUAU_CGCAGACUU_GUCGCUAUCGUGACAAUUAGUUAGCAGCAU .......................((.((((((.........(((((................................))))).....(((((....)))))..)))))).))..... (-11.01 = -11.39 + 0.38)


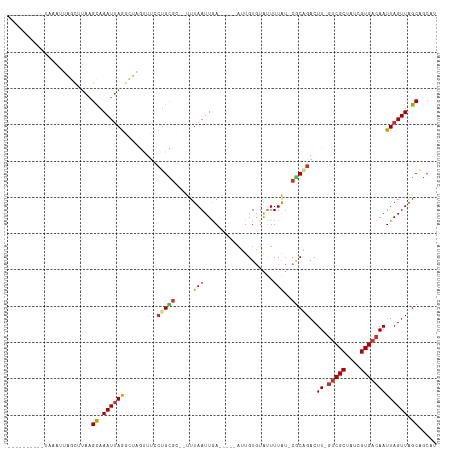
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:22:03 2011