| Sequence ID | dm3.chr3R |
|---|---|
| Location | 4,763,455 – 4,763,554 |
| Length | 99 |
| Max. P | 0.998152 |

| Location | 4,763,455 – 4,763,554 |
|---|---|
| Length | 99 |
| Sequences | 9 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 63.57 |
| Shannon entropy | 0.70325 |
| G+C content | 0.31761 |
| Mean single sequence MFE | -19.18 |
| Consensus MFE | -5.57 |
| Energy contribution | -5.36 |
| Covariance contribution | -0.21 |
| Combinations/Pair | 1.64 |
| Mean z-score | -3.28 |
| Structure conservation index | 0.29 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 3.27 |
| SVM RNA-class probability | 0.998152 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
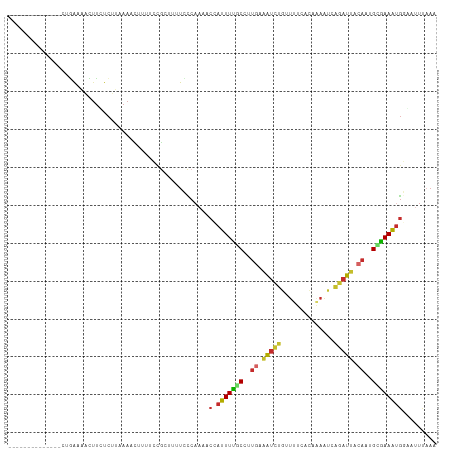
>dm3.chr3R 4763455 99 + 27905053 --------------CUGAAAACUUCUCUUCAAACUUUUCCGCUUUUCCCAAAACCAUUUUGCUUUGAAAUCUGUUUUUACCAAAUCAGAUUAAACUGCAAAAUGGAACUUAAA --------------.((((........))))......................(((((((((.....((((((.(((....))).)))))).....)))))))))........ ( -18.40, z-score = -3.98, R) >droMoj3.scaffold_6540 20215603 86 + 34148556 --------------UUUGAAAAUAUUCUUAAAAUUCUUUU----UUCUCUUCAC-AUUUUUCAUUGUGUUUU--CCGCAC------AGAGUUCAAUGGAAAAUGCAACUCUAA --------------...(((((................))----)))......(-(((((((((((..((((--......------))))..))))))))))))......... ( -12.89, z-score = -1.48, R) >droWil1.scaffold_181089 5959233 108 - 12369635 UAAAAAUUUUUCGAUGAGUUGUAUAUUCAAAAGAUUUCCUUUUUCUUGCAACAUAAUUUUUCCCUAUCAAUUAGUUUUUCUUAGU--CAUCUCUUUGAGAAGACUUUUUC--- .................((((((.....(((((.....)))))...))))))..............................(((--(.((((...)))).)))).....--- ( -13.90, z-score = -0.25, R) >droPer1.super_3 1598147 97 - 7375914 --------------CGGAAAACUUGUCUAAAAAUGCUCUUGAAAUUUCCAACAC-AUUUCCCAUUGCAAUCUAAUCCCAUUGAAU-GGAUUGCAAAGCGAAAUGAUAUUCAAA --------------.(((((.....((.............))..)))))....(-(((((.(.((((((((((.((.....)).)-))))))))).).))))))......... ( -22.42, z-score = -3.31, R) >dp4.chr2 7791191 97 - 30794189 --------------CGGAAAACUUGUCUAAAAAUGCUCUUGAAAUUUCCAACAC-AUUUCCCAUUGCAAUCUAAUCCCAUUGAAU-GGAUUGCAAAGCGAAAUGAUAUUCAAA --------------.(((((.....((.............))..)))))....(-(((((.(.((((((((((.((.....)).)-))))))))).).))))))......... ( -22.42, z-score = -3.31, R) >droEre2.scaffold_4770 906324 98 - 17746568 --------------CUGAAAACUUUUCCUCAAACUUUUCCGCUUUUCCCAAA-CCAUUUUGCCUUGAGAUCUGUUUUCACAAAAUCAGAUUACAAUGCGAAAUGGAAUUUAAA --------------..(((((.(((.....))).))))).............-(((((((((.(((.((((((.(((....))).)))))).))).)))))))))........ ( -21.70, z-score = -3.98, R) >droYak2.chr3R 8807549 99 + 28832112 --------------CUUAAAACUUUUCCUAAAACUUUUCCGCUUUUCCCAAAACCAUUUUGCCUUGAGAUCUGCUUUCACCAAAUCAGAUUACAAUGCGAAAUGGAAUUUAAA --------------.......................................(((((((((.(((.((((((.(((....))).)))))).))).)))))))))........ ( -21.00, z-score = -4.64, R) >droSec1.super_0 4004419 99 - 21120651 --------------CUGAAAACUUCUCUUCAAACUUUUCCGCUUUUCCCAAAACCAUUUUGCCUUGAAAUCUGUUUUUACAAAAUCAGAUUACACUGCGAAAUGGAAUUUAAA --------------.((((........))))......................(((((((((..((.((((((.(((....))).)))))).))..)))))))))........ ( -20.00, z-score = -4.29, R) >droSim1.chr3R 10964652 99 - 27517382 --------------CUGAAAACUUCUCUUCAAACUUUUCCGCUUUUCCCAAAACCAUUUUGCCUUGAAAUCUGUUUUUACAAAAUCAGAUUACACCGCGAAAUGGAAUUUAAA --------------.((((........))))......................(((((((((..((.((((((.(((....))).)))))).))..)))))))))........ ( -19.90, z-score = -4.26, R) >consensus ______________CUGAAAACUUCUCUUAAAACUUUUCCGCUUUUCCCAAAACCAUUUUGCCUUGAAAUCUGUUUUCACAAAAUCAGAUUACAAUGCGAAAUGGAAUUUAAA .......................................................(((((((..((.(((((..............))))).))..))))))).......... ( -5.57 = -5.36 + -0.21)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:59:31 2011