
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 4,271,164 – 4,271,256 |
| Length | 92 |
| Max. P | 0.570394 |

| Location | 4,271,164 – 4,271,256 |
|---|---|
| Length | 92 |
| Sequences | 9 |
| Columns | 106 |
| Reading direction | reverse |
| Mean pairwise identity | 83.72 |
| Shannon entropy | 0.31043 |
| G+C content | 0.50738 |
| Mean single sequence MFE | -21.78 |
| Consensus MFE | -10.93 |
| Energy contribution | -10.84 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.29 |
| Structure conservation index | 0.50 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.16 |
| SVM RNA-class probability | 0.570394 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 4271164 92 - 27905053 AGCGGGACCCCCCGCACACACACAUGCACGACAAAGCAGAAUAAUAAAUUUCAAUUCAGACGAC--------GACAUGACGUCUCUGCCUCACGUCGACG------ .(((((....))))).............((((......(((........)))....((((.(((--------(......))))))))......))))...------ ( -25.00, z-score = -3.57, R) >droSim1.chr3R 10512962 90 + 27517382 AGCGGGACCCCCCGC--ACACACAUGCACGACAAAGCAGAAUAAUAAAUUUCAAUUCAGACGAC--------GACAUGACGUCUCUGCCUCACGUCGACG------ .(((((....)))))--...........((((......(((........)))....((((.(((--------(......))))))))......))))...------ ( -25.00, z-score = -3.57, R) >droSec1.super_0 3552597 90 + 21120651 AGCGGGACCCCCCGC--ACACACAUGCACGACAAAGCAGAAUAAUAAAUUUCAAUUCAGACGAC--------GACACGACGUCUCUGCCUCACGUCGACG------ .(((((....)))))--...........((((......(((........)))....((((.(((--------(......))))))))......))))...------ ( -25.00, z-score = -3.68, R) >droYak2.chr3R 8341699 90 - 28832112 AGCGGGACCCCCCGC--ACACACAUGCACGACAAAGCAGAAUAAUAAAUUUCAAUUCAGACGAC--------GACAUGACGUCUCUGCCUCACGUCGACG------ .(((((....)))))--...........((((......(((........)))....((((.(((--------(......))))))))......))))...------ ( -25.00, z-score = -3.57, R) >droEre2.scaffold_4770 440744 88 + 17746568 AGCGGGACCCCGC----ACACACAUGCACGACAAAGCAGAAUAAUAAAUUUCAAUUCAGACGAC--------GACAUGACGUCUCUGCCUCACGUCGACG------ .((((....))))----...........((((......(((........)))....((((.(((--------(......))))))))......))))...------ ( -23.10, z-score = -2.80, R) >droAna3.scaffold_13340 19565274 79 + 23697760 --------CUCC-----UCACACACGCACGACAAAGCAGAAUAAUAAAUUUCAAUUCAGACGAC--------GACAUGACGUCUCUGCCUCACGUCGACG------ --------....-----...........((((......(((........)))....((((.(((--------(......))))))))......))))...------ ( -14.80, z-score = -1.55, R) >droPer1.super_3 6003381 87 - 7375914 ------ACCCAC-----ACACACACGCACGACAAAGCAGAAUAAUAAAUUUCAAUUCAGACGAC--------GACAUGACGUCUCUGCCUCACGUCGACCGUCGAC ------......-----......(((..((((......(((........)))....((((.(((--------(......))))))))......))))..))).... ( -15.90, z-score = -1.31, R) >droWil1.scaffold_181089 832627 88 - 12369635 ---AGAGUGUUUC----ACACACACGCACGACAAAGCAGAAUAAUAAAUUUCAAUUCAGACGACG-----AGGACAUGACGUCUCUGCUUCACGUCGACG------ ---...((((...----..)))).....((((.(((((((..........((......)).((((-----.........)))))))))))...))))...------ ( -18.80, z-score = -0.60, R) >droMoj3.scaffold_6540 3327392 97 - 34148556 CUCAGAAGUGCCUG---GCACACACGCACGACAAAGCAGAAUAAUAAAUUUCAAUUCAGACGACGAGAGCAGCGCAGAGCGUCUCUGCCGCACGUUGACG------ ....(((((((.((---.....)).)))).........(((........)))..)))...(((((...((.(.((((((...))))))))).)))))...------ ( -23.40, z-score = 0.04, R) >consensus AGCGGGACCCCCCG___ACACACAUGCACGACAAAGCAGAAUAAUAAAUUUCAAUUCAGACGAC________GACAUGACGUCUCUGCCUCACGUCGACG______ ............................((((...(((((..........((......))............(((.....)))))))).....))))......... (-10.93 = -10.84 + -0.08)


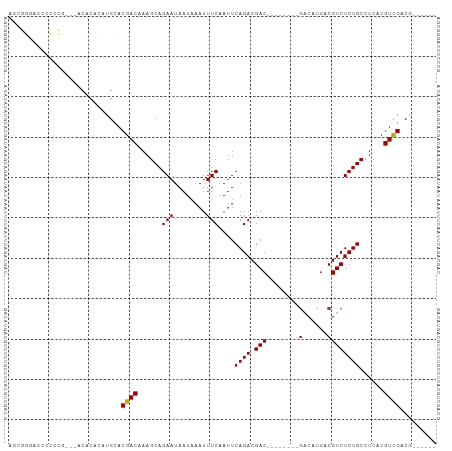
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:58:22 2011