
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 4,022,559 – 4,022,649 |
| Length | 90 |
| Max. P | 0.533285 |

| Location | 4,022,559 – 4,022,649 |
|---|---|
| Length | 90 |
| Sequences | 10 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 77.29 |
| Shannon entropy | 0.45204 |
| G+C content | 0.31400 |
| Mean single sequence MFE | -11.70 |
| Consensus MFE | -5.58 |
| Energy contribution | -5.50 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.48 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.08 |
| SVM RNA-class probability | 0.533285 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
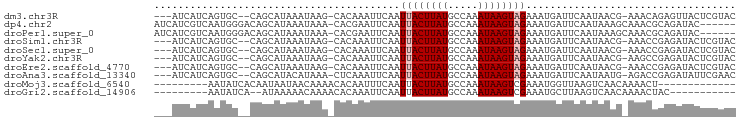
>dm3.chr3R 4022559 90 + 27905053 ---AUCAUCAGUGC--CAGCAUAAAUAAG-CACAAAUUCAAUUACUUAUGCCAAAUAAGUAGAAAUGAUUCAAUAACG-AAACAGAGUUACUCGUAC ---.......((((--............)-))).(((((..((((((((.....))))))))...((.(((......)-)).)))))))........ ( -13.90, z-score = -2.01, R) >dp4.chr2 17748202 90 - 30794189 AUCAUCGUCAAUGGGACAGCAUAAAUAAA-CACGAAUUCAAUUACUUAUGCCAAAUAAGUAGAAAUGAUUCAAUAAAGCAAACGCAGAUAC------ ...((((((.....)))............-...(((((...((((((((.....))))))))....)))))......((....)).)))..------ ( -13.40, z-score = -1.73, R) >droPer1.super_0 7409061 90 + 11822988 AUCAUCGUCAAUGGGACAGCAUAAAUAAA-CACGAAUUCAAUUACUUAUGCCAAAUAAGUAGAAAUGAUUCAAUAAAGCAAACGCAGAUAC------ ...((((((.....)))............-...(((((...((((((((.....))))))))....)))))......((....)).)))..------ ( -13.40, z-score = -1.73, R) >droSim1.chr3R 10282474 90 - 27517382 ---AUCAUCAGUGC--CAGCAUAAAUAAG-CACAAAUUCAAUUACUUAUGCCAAAUAAGUAGAAAUGAUUCAAUAACG-AAACCGAGAUACUCGUAC ---(((((..((((--............)-)))........((((((((.....))))))))..))))).........-....((((...))))... ( -13.50, z-score = -2.04, R) >droSec1.super_0 3314470 90 - 21120651 ---AUCAUCAGUGC--CAGCAUAAAUAAG-CACAAAUUCAAUUACUUAUGCCAAAUAAGUAGAAAUGAUUCAAUAACG-AAACCGAGAUACUCGUAC ---(((((..((((--............)-)))........((((((((.....))))))))..))))).........-....((((...))))... ( -13.50, z-score = -2.04, R) >droYak2.chr3R 8096708 90 + 28832112 ---AUCAUCAGUGC--CAGCAUAAAUAAG-CACAAAUUCAAUUACUUAUGCCAAAUAAGUAGAAAUGAUUCAAUAACG-AAGCCGAGAUACUCGUAC ---(((((..((((--............)-)))........((((((((.....))))))))..))))).........-....((((...))))... ( -13.50, z-score = -1.60, R) >droEre2.scaffold_4770 190957 90 - 17746568 ---AUCAUCAGUGC--CAGCAUAAAUAAG-CACAAAUUCAAUUACUUAUGCCAAAUAAGUAGAAAUGAUUCAAUAACG-AAACCGAGAUACUCGUAC ---(((((..((((--............)-)))........((((((((.....))))))))..))))).........-....((((...))))... ( -13.50, z-score = -2.04, R) >droAna3.scaffold_13340 21476596 90 + 23697760 ---AUCAUCAGUGC--CAGCAUACAUAAA-CUCAAAUUCAAUUACUUAUGCCAAAUAAGUAGAAAUGAUUCAAUAAUG-AGACCGAGAUAUUCGAAC ---(((.((.(((.--...))).......-((((........(((((((.....)))))))(((....))).....))-))...)))))........ ( -11.90, z-score = -0.98, R) >droMoj3.scaffold_6540 24840053 75 - 34148556 ---------AAUAUCACAAUAAUAACAAAACACAAUUUCAAUUACUUAUGCCAAAUAAGUCGAAAUGGUUAAGUCAACAAAACU------------- ---------...................(((...(((((....((((((.....)))))).))))).)))..............------------- ( -7.10, z-score = -1.72, R) >droGri2.scaffold_14906 6765498 75 - 14172833 ---------AAUAUCA--AUAAAAACAAAACACAAAUUCAAUUACUUAUGCCAAAUAAGUCGAAAUGCUUAAGUCAACAAAACUAC----------- ---------.......--..................(((....((((((.....)))))).)))......................----------- ( -3.30, z-score = -0.45, R) >consensus ___AUCAUCAGUGC__CAGCAUAAAUAAA_CACAAAUUCAAUUACUUAUGCCAAAUAAGUAGAAAUGAUUCAAUAACG_AAACCGAGAUACUCGUAC ..........((((....))))...................((((((((.....))))))))................................... ( -5.58 = -5.50 + -0.08)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:57:48 2011