
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 3,983,216 – 3,983,344 |
| Length | 128 |
| Max. P | 0.779767 |

| Location | 3,983,216 – 3,983,308 |
|---|---|
| Length | 92 |
| Sequences | 10 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 60.12 |
| Shannon entropy | 0.80727 |
| G+C content | 0.45405 |
| Mean single sequence MFE | -21.52 |
| Consensus MFE | -5.38 |
| Energy contribution | -5.46 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.67 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.25 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.67 |
| SVM RNA-class probability | 0.779767 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
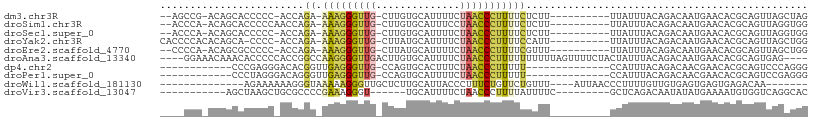
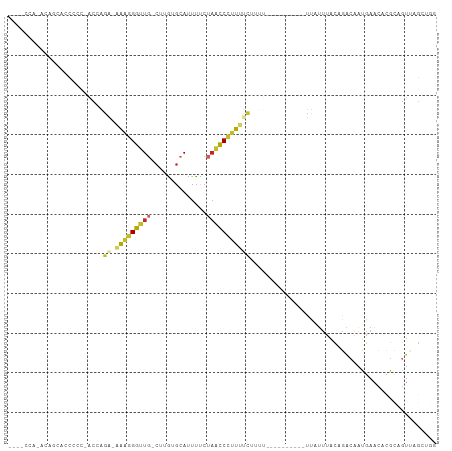
>dm3.chr3R 3983216 92 - 27905053 --AGCCG-ACAGCACCCCC-ACCAGA-AAAGGGUUG-CUUGUGCAUUUUCUAACCCUUUUCUCUU----------UUAUUUACAGACAAUGAACACGCAGUUAGCUAG --(((..-((.((......-...(((-(((((((((-(....))).......))))))))))..(----------(((((.......))))))...)).))..))).. ( -22.41, z-score = -2.89, R) >droSim1.chr3R 10241801 93 + 27517382 --ACCCA-ACAGCACCCCCAACCAGA-AAAGGGUUG-CUUGUGCAUUUCCUAACCCUUUUCUCUU----------UUAUUUACAGACAAUGAACACGCAGUUAGGUGG --(((.(-((.((..........(((-(((((((((-(....))).......))))))))))..(----------(((((.......))))))...)).))).))).. ( -21.31, z-score = -1.65, R) >droSec1.super_0 3275390 92 + 21120651 --ACCCA-ACAGCACCCCC-ACCAGA-AAAGGGUUG-CUUGUGCAUUUUCUAACCCUUUUCUCUU----------UUAUUUACAGACAAUGAACACGCAGUUAGGUGG --.....-.........((-((((((-(((((((((-(....))).......))))))))))..(----------(((((.......))))))..........))))) ( -22.41, z-score = -2.05, R) >droYak2.chr3R 8055921 94 - 28832112 CACCCCACACAGCA-CCCC-ACCAGA-AAAGGGUUG-CUUAUGCAUUUUCUAACCCUUUUCCAUU----------UUAUUUACAGACAAUGAACACGCAGUUAGCUGG .........((((.-....-....((-(((((((((-(....))).......)))))))))...(----------(((((.......))))))..........)))). ( -19.61, z-score = -2.75, R) >droEre2.scaffold_4770 149045 92 + 17746568 --CCCCA-ACAGCGCCCCC-ACCAGA-AAAGGGUUG-CUUAUGCAUUUUCUAACCCUUUUCGUUU----------UUAUUUACAGACAAUGAACACGCAGUUAGCUGG --.....-.((((......-....((-(((((((((-(....))).......)))))))))((.(----------(((((.......)))))).)).......)))). ( -22.81, z-score = -2.95, R) >droAna3.scaffold_13340 21438117 100 - 23697760 ----GGAAACAAACACCCCCACCGGCCAAGGGGUUGACUUGUGCAUUUUCUAACCCUUUUUUUUUUAGUUUUCUACUAUUUACAGACAAUGAACACGCAGUGAG---- ----(....)...((.((((.........)))).))(((((((((((.(((..............((((.....)))).....))).))))..)))).)))...---- ( -15.21, z-score = 0.90, R) >dp4.chr2 17706459 81 + 30794189 ------------CCCGAGGGACACGGUUGAGGGUUG-CCAGUGCACUUUCUAACCCUUUUU--------------CCAUUUACAGACAACGAACACGCAGUCCCAGGG ------------(((..(((((..((..((((((((-..((....))...))))))))...--------------))............((....))..))))).))) ( -25.40, z-score = -1.96, R) >droPer1.super_0 7366594 81 - 11822988 ------------CCCUAGGGACAGGGUUGAGGGUUG-CCAGUGCAUUUUCUAACCCUUUUU--------------CCAUUUACAGACAACGAACACGCAGUCCGAGGG ------------((((..((((.(((..((((((((-..((.....))..))))))))..)--------------))............((....))..)))).)))) ( -24.70, z-score = -1.56, R) >droWil1.scaffold_181130 14080673 83 - 16660200 --------------AGAAAAAAGGGUAAAAAGGGUUGCUCUUGCAUUACCCUUUCUGUUCUGUUU----AUUAACCCUUUUGUUGUGAGUGAGUGAGACAA------- --------------.......((..((.((((((((((....)))..))))))).))..))((((----....((((((.......))).).)).))))..------- ( -18.60, z-score = -0.82, R) >droVir3.scaffold_13047 10004287 82 - 19223366 -----------AGCUAAGCUGCGCCCCGAAAGGGU------UGCAUUUUCUAACCCUUUUAUUUUC---------GCUCAGACAAUAUAUGAAAAUGUGGUCAGGCAC -----------.((......))(((..((((((((------((.......))))))))))......---------.....(((..(((((....)))))))).))).. ( -22.70, z-score = -1.88, R) >consensus ____CCA_ACAGCACCCCC_ACCAGA_AAAGGGUUG_CUUGUGCAUUUUCUAACCCUUUUCUUUU__________UUAUUUACAGACAAUGAACACGCAGUUAGCUGG ...........................(((((((((..............)))))))))................................................. ( -5.38 = -5.46 + 0.08)

| Location | 3,983,244 – 3,983,344 |
|---|---|
| Length | 100 |
| Sequences | 11 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 62.79 |
| Shannon entropy | 0.75871 |
| G+C content | 0.42981 |
| Mean single sequence MFE | -23.96 |
| Consensus MFE | -5.64 |
| Energy contribution | -5.64 |
| Covariance contribution | -0.01 |
| Combinations/Pair | 1.36 |
| Mean z-score | -1.86 |
| Structure conservation index | 0.24 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.691034 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
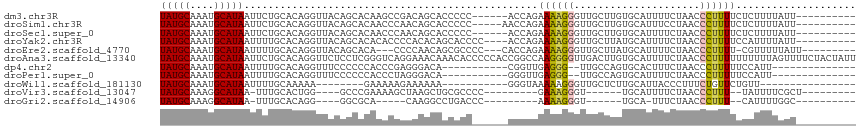
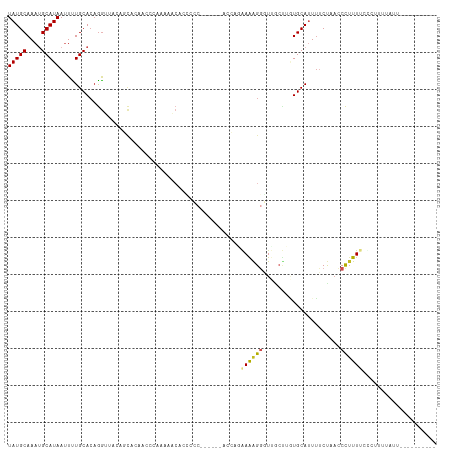
>dm3.chr3R 3983244 100 - 27905053 UAUGCAAAUGCAUAAUUCUGCACAGGUUACAGCACAAGCCGACAGCACCCCC------ACCAGAAAAGGGUUGCUUGUGCAUUUUCUAACCCUUUUCUCUUUUAUU---------- (((((....)))))....(((...((((........))))....))).....------...(((((((((((((....))).......))))))))))........---------- ( -24.01, z-score = -2.02, R) >droSim1.chr3R 10241829 101 + 27517382 UAUGCAAAUGCAUAAUUCUGCACAGGUUACAGCACAACCCAACAGCACCCCC-----AACCAGAAAAGGGUUGCUUGUGCAUUUCCUAACCCUUUUCUCUUUUAUU---------- ..(((...((((......))))..((((.......)))).....))).....-----....(((((((((((((....))).......))))))))))........---------- ( -22.01, z-score = -1.92, R) >droSec1.super_0 3275418 100 + 21120651 UAUGCAAAUGCAUAAUUCUGCACAGGUUACAGCACAACCCAACAGCACCCCC------ACCAGAAAAGGGUUGCUUGUGCAUUUUCUAACCCUUUUCUCUUUUAUU---------- ..(((...((((......))))..((((.......)))).....))).....------...(((((((((((((....))).......))))))))))........---------- ( -22.01, z-score = -1.96, R) >droYak2.chr3R 8055949 102 - 28832112 UAUGCAAAUGCAUAAUUUUGCACAGGUUACAGCACACACCCCACACAGCACCCC----ACCAGAAAAGGGUUGCUUAUGCAUUUUCUAACCCUUUUCCAUUUUAUU---------- ..((((((........))))))..(((....((..............)).....----))).((((((((((((....))).......))))))))).........---------- ( -20.35, z-score = -2.12, R) >droEre2.scaffold_4770 149073 100 + 17746568 UAUGCAAAUGCAUAAUUUUGCACAGGUUACAGCACA---CCCCAACAGCGCCCC---CACCAGAAAAGGGUUGCUUAUGCAUUUUCUAACCCUUUU-CGUUUUUAUU--------- ..((((((........))))))..(((....((.(.---........).))...---.))).((((((((((((....))).......))))))))-).........--------- ( -22.71, z-score = -2.24, R) >droAna3.scaffold_13340 21438141 116 - 23697760 UAUGCAAAUGCAUAAUUCUGCACAGGUUCUCCUCGGGUCAGGAAACAAACACCCCCACCGGCCAAGGGGUUGACUUGUGCAUUUUCUAACCCUUUUUUUUUUAGUUUUCUACUAUU (((((....)))))....((((((((((..((((.((((.((...............))))))..))))..))))))))))....................((((.....)))).. ( -31.56, z-score = -2.38, R) >dp4.chr2 17706487 89 + 30794189 UAUGCAAAUGCAUAAUUUUGCACAGGUUUCCCCCCACCCGAGGGACA-----------CGGUUGAGGG--UUGCCAGUGCACUUUCUAACCCUUUUUCCAUU-------------- (((((....)))))....(((((.(((..(((..((.(((.......-----------))).)).)))--..))).))))).....................-------------- ( -25.60, z-score = -1.72, R) >droPer1.super_0 7366622 89 - 11822988 UAUGCAAAUGCAUAAUUUUGCACAGGUUUCCCCCCACCCUAGGGACA-----------GGGUUGAGGG--UUGCCAGUGCAUUUUCUAACCCUUUUUCCAUU-------------- (((((....)))))....(((((.(((..(((.(.(((((......)-----------)))).).)))--..))).))))).....................-------------- ( -29.90, z-score = -2.52, R) >droWil1.scaffold_181130 14080706 81 - 16660200 UAUGCAAAUGCAUAAUUUUGCAAAAA--------GAAAAAGAAAAAA-----------GGGUAAAAAGGGUUGCUCUUGCAUUACCCUUUCUGUUCUGUU---------------- ..((((((........))))))....--------............(-----------(..((.((((((((((....)))..))))))).))..))...---------------- ( -21.30, z-score = -2.87, R) >droVir3.scaffold_13047 10004317 85 - 19223366 UAUGCAAAGGCAUAA-UUUGCACUGG----GCCCGAAAAGCUAAGCUGCGCCCC---------GAAAGGGU------UGCAUUUUCUAACCCUUU--UAUUUUCGCU--------- ...((..(((((...-..))).))..----)).((((((((......)).....---------((((((((------((.......)))))))))--).))))))..--------- ( -23.60, z-score = -0.64, R) >droGri2.scaffold_14906 6721712 78 + 14172833 UAUGCAAAGGCAUAA-UUUGCACAGG----GGCGCA-----CAAGGCCUGACCC---------AAAAGGGU------UGCA-UUUCUAACCCUUU--CAUUUUGGC---------- ..((((((.......-))))))...(----(((...-----....))))...((---------((((((((------((..-....)))))))..--...))))).---------- ( -20.50, z-score = -0.11, R) >consensus UAUGCAAAUGCAUAAUUUUGCACAGGUUACAGCACAACCCAAAAACACCCCC______ACCAGAAAAGGGUUGCUUGUGCAUUUUCUAACCCUUUUCCCUUUUAUU__________ (((((....))))).................................................((((((.....................)))))).................... ( -5.64 = -5.64 + -0.01)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:57:40 2011