
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 3,903,355 – 3,903,418 |
| Length | 63 |
| Max. P | 0.896309 |

| Location | 3,903,355 – 3,903,418 |
|---|---|
| Length | 63 |
| Sequences | 9 |
| Columns | 71 |
| Reading direction | forward |
| Mean pairwise identity | 80.51 |
| Shannon entropy | 0.37989 |
| G+C content | 0.53146 |
| Mean single sequence MFE | -18.59 |
| Consensus MFE | -12.91 |
| Energy contribution | -13.00 |
| Covariance contribution | 0.09 |
| Combinations/Pair | 1.07 |
| Mean z-score | -1.74 |
| Structure conservation index | 0.69 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.13 |
| SVM RNA-class probability | 0.896309 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
>dm3.chr3R 3903355 63 + 27905053 UCAUUUGGCAUGCAGCAAACGUGUCAAGGACAAGUCGGAUGCAGCCACCGACU-----CGACACAGAA--- ((.(((((((((.......)))))))))))(.((((((.((....))))))))-----.)........--- ( -16.90, z-score = -1.10, R) >droEre2.scaffold_4770 78585 63 - 17746568 UCAUUUGGCAUGCAGCAAACGUGUCAAGGACAAGUCUGAUGCAGCCACCGACU-----CGACACACAA--- .....((((.((((.((.((.((((...)))).)).)).))))))))......-----..........--- ( -16.30, z-score = -1.00, R) >droYak2.chr3R 7984632 63 + 28832112 UCAUUUGGCAUGCAGCAAACGUGUCAAGGACAAGUCGGAUGCAACCACCGACU-----CGACACACAG--- ((.(((((((((.......)))))))))))(.((((((.((....))))))))-----.)........--- ( -16.90, z-score = -1.71, R) >droSec1.super_0 3207120 63 - 21120651 UCAUUUGGCAUGCAGCAAACGUGUCAAGGACAAGUCGGAUGCAGCCACCGACU-----CGACACACAA--- ((.(((((((((.......)))))))))))(.((((((.((....))))))))-----.)........--- ( -16.90, z-score = -1.21, R) >droSim1.chr3R 10171262 63 - 27517382 UCAUUUGGCAUGCAGCAAACGUGUCAAGGACAAGUCGGAUGCAGCCACCGACU-----CGACACACAA--- ((.(((((((((.......)))))))))))(.((((((.((....))))))))-----.)........--- ( -16.90, z-score = -1.21, R) >droPer1.super_0 209690 50 + 11822988 UCAUUUGGCAUGCAGCAGGCGUGUCAAGGACAAGUCUGAUGCAGCCAUUG--------------------- .....((((.((((.(((((.((((...)))).))))).))))))))...--------------------- ( -22.00, z-score = -3.63, R) >dp4.chr2 6385216 50 + 30794189 UCAUUUGGCAUGCAGCAGGCGUGUCAAGGACAAGUCUGAUGCAGCCAUUG--------------------- .....((((.((((.(((((.((((...)))).))))).))))))))...--------------------- ( -22.00, z-score = -3.63, R) >droAna3.scaffold_13340 15173820 71 - 23697760 UCAUUGGGCAUGCAGCAAGCGUGUCAAGGACAUGUCUGAUGCAGCCACCGACCACCGACCACCCACCACCA ......(((.((((.((..((((((...))))))..)).)))))))......................... ( -20.30, z-score = -0.93, R) >droVir3.scaffold_13047 9928926 62 + 19223366 UCAUUUGGCAUGCAAGAAGCGUGCCAAGGACAAGUCAGCUGCAGCCAC-GCCU-----CCAAACGCAA--- ....(((((((((.....)))))))))(((...((..((....)).))-...)-----))........--- ( -19.10, z-score = -1.24, R) >consensus UCAUUUGGCAUGCAGCAAACGUGUCAAGGACAAGUCGGAUGCAGCCACCGACU_____CGACACACAA___ .....((((.((((....((.((((...)))).))....))))))))........................ (-12.91 = -13.00 + 0.09)


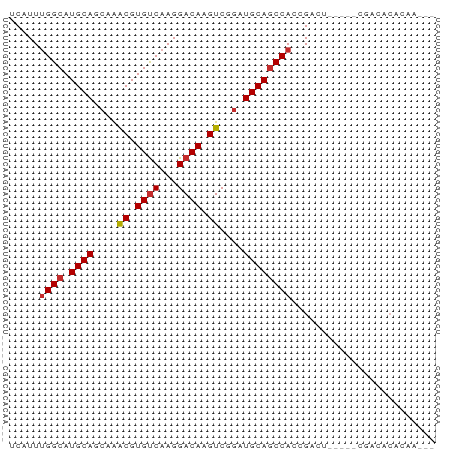
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:57:23 2011