
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 3,476,976 – 3,477,047 |
| Length | 71 |
| Max. P | 0.500000 |

| Location | 3,476,976 – 3,477,047 |
|---|---|
| Length | 71 |
| Sequences | 4 |
| Columns | 84 |
| Reading direction | forward |
| Mean pairwise identity | 63.16 |
| Shannon entropy | 0.57108 |
| G+C content | 0.29958 |
| Mean single sequence MFE | -11.80 |
| Consensus MFE | -6.70 |
| Energy contribution | -8.21 |
| Covariance contribution | 1.50 |
| Combinations/Pair | 1.28 |
| Mean z-score | -0.62 |
| Structure conservation index | 0.57 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
>dm3.chr3R 3476976 71 + 27905053 -------UUUGAAAAUGUUACAGAUUCCACGCUUAUUUAUGGCGAA------UAUAAUUAUAAUUAAAUGUGGAGGCAUGAGGC -------.........((..((..(((((((.(((.((((((....------.....)))))).))).)))))))...))..)) ( -12.60, z-score = -1.00, R) >droSec1.super_6 3568416 65 + 4358794 -------UUUGCAAAUUUUAGCACUUCCAUGCUUAUUUAUAGCGAA------UAUAAGUAUAAUUAAAUAUGGAAGCA------ -------..(((........)))((((((((.(((.(((((.(...------.....)))))).))).))))))))..------ ( -14.30, z-score = -1.71, R) >droYak2.chr3R_random 1085817 71 + 3277457 -----------CUAGACUUAGCGCUUAGAUAUUACUUUAAAGCGAACCACUUUAUAAGUAUUAUUAAAUAUUGAGGCAGAAA-- -----------.........((.((((((((.((((((((((.......))))).))))).))))......)))))).....-- ( -10.30, z-score = -0.58, R) >droEre2.scaffold_4770 3736675 82 + 17746568 CCCGCAAAUUGCAAUUCUAA--GAUUGGAUGUUAAUUUAUGGCGAACAACUUUAUAAGUAUAAUUAAAUAUUGAGGCAGAAGGA .((((.....))..((((..--..(((..(((((.....)))))..)))(((.(((..((....))..))).)))..)))))). ( -10.00, z-score = 0.79, R) >consensus _______UUUGCAAAUCUUAGCGAUUCCAUGCUAAUUUAUAGCGAA______UAUAAGUAUAAUUAAAUAUGGAGGCAGAAG__ ......................(((((((((((((.(((((.................))))).)))))))))))))....... ( -6.70 = -8.21 + 1.50)


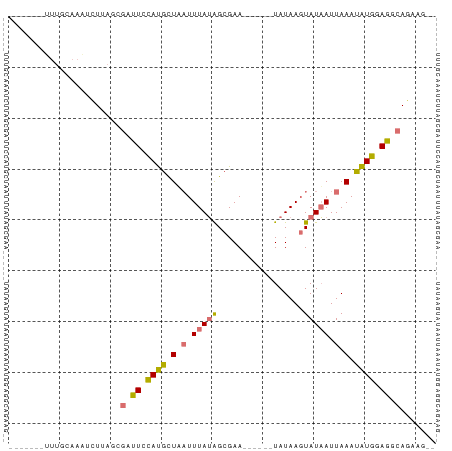
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:56:36 2011