
| Sequence ID | dm3.chr2LHet |
|---|---|
| Location | 316,478 – 316,591 |
| Length | 113 |
| Max. P | 0.884508 |

| Location | 316,478 – 316,591 |
|---|---|
| Length | 113 |
| Sequences | 3 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 59.31 |
| Shannon entropy | 0.55605 |
| G+C content | 0.54277 |
| Mean single sequence MFE | -38.40 |
| Consensus MFE | -18.87 |
| Energy contribution | -15.23 |
| Covariance contribution | -3.64 |
| Combinations/Pair | 1.55 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.06 |
| SVM RNA-class probability | 0.884508 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2LHet 316478 113 + 368872 ----UUUCUGCCCGCACAGGGGCAACCUGCA-UAGCCGCGAGCUGGCCGGCAAGUUCCUGAACUGUAUUACGCAGGUCGGCCUCCCUGCGUUGACCUUCUAGGGCUUGCUCUUGUAAC ----.....(((.((...(((....)))((.-.....))..)).))).(((((((((..(((..((...(((((((........)))))))..)).)))..)))))))))........ ( -40.40, z-score = -0.37, R) >dp4.Unknown_group_92 21448 113 + 26366 CUUGUGCUCAACUGAGCAAUCGCGGCUUGUAAUUACAGUAACCUGGACU-CAAACCACGAGGCCUGAUUUUGCAGUGCGGCUCCUAUA-GCCGA---UGCGAAAGUUGCCCGAUCCAU ....(((((....))))).(((.((.(((......(((....)))....-))).)).)))(((....(((((((...(((((.....)-)))).---)))))))...)))........ ( -35.40, z-score = -1.80, R) >droPer1.super_143 87038 113 - 108547 CUUGUGCUCAACUGAGCAAUCGCGGCUUGUAAUUACAGUAACCUGGUCU-CAAACCGCGAGGCCUGAUUUUGCAGUGCGGCCUCUAUA-GCCGA---UGCGAAAGUUGCCCGAUCCAU ....(((((....))))).((((((.(((......(((....)))....-))).))))))(((....(((((((...((((.......-)))).---)))))))...)))........ ( -39.40, z-score = -2.49, R) >consensus CUUGUGCUCAACUGAGCAAUCGCGGCUUGUAAUUACAGUAACCUGGACU_CAAACCACGAGGCCUGAUUUUGCAGUGCGGCCUCUAUA_GCCGA___UGCGAAAGUUGCCCGAUCCAU ....(((((....))))).(((.(....((....)).(((((............((....)).....(((((((...((((........))))....))))))))))))))))..... (-18.87 = -15.23 + -3.64)


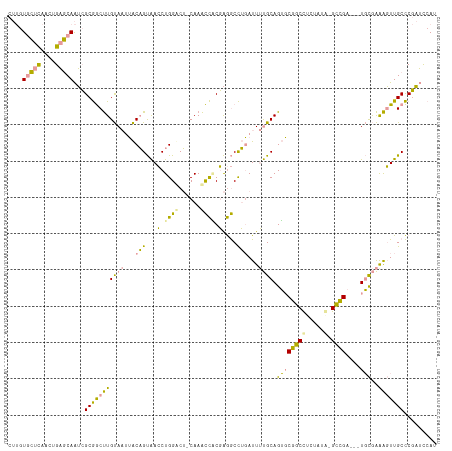
| Location | 316,478 – 316,591 |
|---|---|
| Length | 113 |
| Sequences | 3 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 59.31 |
| Shannon entropy | 0.55605 |
| G+C content | 0.54277 |
| Mean single sequence MFE | -39.48 |
| Consensus MFE | -21.10 |
| Energy contribution | -20.22 |
| Covariance contribution | -0.88 |
| Combinations/Pair | 1.35 |
| Mean z-score | -1.29 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.89 |
| SVM RNA-class probability | 0.845792 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2LHet 316478 113 - 368872 GUUACAAGAGCAAGCCCUAGAAGGUCAACGCAGGGAGGCCGACCUGCGUAAUACAGUUCAGGAACUUGCCGGCCAGCUCGCGGCUA-UGCAGGUUGCCCCUGUGCGGGCAGAAA---- .............((((.....(.....)(((.((.(((.((((((((((....((((....)))).((((.(......)))))))-))))))))))).)).))))))).....---- ( -41.90, z-score = -0.42, R) >dp4.Unknown_group_92 21448 113 - 26366 AUGGAUCGGGCAACUUUCGCA---UCGGC-UAUAGGAGCCGCACUGCAAAAUCAGGCCUCGUGGUUUG-AGUCCAGGUUACUGUAAUUACAAGCCGCGAUUGCUCAGUUGAGCACAAG ...((((((((...(((.(((---.((((-(.....)))))...))).)))....)))).((((((((-(((.(((....)))..))).))))))))))))((((....))))..... ( -37.60, z-score = -1.53, R) >droPer1.super_143 87038 113 + 108547 AUGGAUCGGGCAACUUUCGCA---UCGGC-UAUAGAGGCCGCACUGCAAAAUCAGGCCUCGCGGUUUG-AGACCAGGUUACUGUAAUUACAAGCCGCGAUUGCUCAGUUGAGCACAAG ((((.((((((.......)).---)))))-))).((((((..............))))))((((((((-.((.(((....)))...)).))))))))...(((((....))))).... ( -38.94, z-score = -1.93, R) >consensus AUGGAUCGGGCAACUUUCGCA___UCGGC_UAUAGAGGCCGCACUGCAAAAUCAGGCCUCGCGGUUUG_AGACCAGGUUACUGUAAUUACAAGCCGCGAUUGCUCAGUUGAGCACAAG .......(((((..............(((........((......))........)))((((((((((.....(((....)))......)))))))))).)))))............. (-21.10 = -20.22 + -0.88)



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:04:51 2011