
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 3,239,835 – 3,239,938 |
| Length | 103 |
| Max. P | 0.500000 |

| Location | 3,239,835 – 3,239,938 |
|---|---|
| Length | 103 |
| Sequences | 9 |
| Columns | 105 |
| Reading direction | forward |
| Mean pairwise identity | 80.14 |
| Shannon entropy | 0.41872 |
| G+C content | 0.46827 |
| Mean single sequence MFE | -32.67 |
| Consensus MFE | -13.83 |
| Energy contribution | -14.24 |
| Covariance contribution | 0.42 |
| Combinations/Pair | 1.41 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.42 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.01 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
>dm3.chr3R 3239835 103 + 27905053 CGACGAGUGUUGCCAAAGUCCUUGUUUGUGGCUGACGAUUUGCCAUUUUAAUUAGGCAAUGUCCUGUCACAAGCAGGCG--GCCAGUGGAUACGGUUAUUAGUUA .(((..((((..(((..(((((((((((((((.((((..(((((..........)))))))))..)))))))))))).)--))...))))))).)))........ ( -36.30, z-score = -2.96, R) >droSim1.chr3R 3284098 103 + 27517382 CGACGAGUGUUGCCAAAGUCCUUGUUUGUGGCUGACGAUUUGCCAUUUUAAUUAGGCAAUGUCCUGUCACAAGCAGGCG--GCCAGUGGAUACGGUUAUUAGUUA .(((..((((..(((..(((((((((((((((.((((..(((((..........)))))))))..)))))))))))).)--))...))))))).)))........ ( -36.30, z-score = -2.96, R) >droSec1.super_6 3331150 103 + 4358794 CGACGAGUGUUGCCAAAGUCCUUGUUUGUGGCUGACGAUUUGCCAUUUUAAUUAGGCAAUGUCCUGUCACAAGCAGGCG--GCCAGUGGAUACGGUUAUUAGUUA .(((..((((..(((..(((((((((((((((.((((..(((((..........)))))))))..)))))))))))).)--))...))))))).)))........ ( -36.30, z-score = -2.96, R) >droYak2.chr3R_random 939479 103 + 3277457 CGACGAGUGUUGCCAAAGUCCUUGUUUGUGGCUGAUGAUUUGCCAUUUUAAUUAGGCAAUGUCCUGUCACAAGCAGACG--GCCAGUAGAUACGGUUAUUAGUUA .(((..((((((((...(((..((((((((((.(((...(((((..........))))).)))..))))))))))))))--)).....))))).)))........ ( -29.70, z-score = -1.80, R) >droEre2.scaffold_4770 3499444 103 + 17746568 CGACGAGUUUUGGCAAAGUCCUUGUUUGUGGCUAAUGAUUUGCCAUUUUAAUUAGGCAAUGUCCUGUCACAAGCAGGCG--GCUAGUGGAUACGGUUAUUAGUUA .(((..((.((.((..((((((((((((((((....((((((((..........))))).)))..)))))))))))).)--))).)).)).)).)))........ ( -32.70, z-score = -2.48, R) >droAna3.scaffold_13340 3026662 101 + 23697760 CGACGAGUGUUGCCGGAGUCCUUGUU-GUGGCUGACGAUUUGCCAUUUUAAUUAGGCAAUGUCCUGUCACAAGCAGCCG--GUCU-CCGACACGGUUAUGAGCUA .(((..((((...(((((.(((((((-(((((.((((..(((((..........)))))))))..))))).)))))..)--).))-))))))).)))........ ( -34.90, z-score = -1.97, R) >dp4.chr2 26632784 104 + 30794189 UGACCAGUGUUGCCAACGUUCUUGUU-GUGGCUGACGAUUUGCCAUUUUAAUUAGGCAAUGUCCUGUCAAAAGCAGCCCCUGGCUGUGGCUCUGGUUAUGGGCAG (((((((....(((((((....))))-.((((.((((..(((((..........)))))))))..))))...((((((...))))))))).)))))))....... ( -34.50, z-score = -1.20, R) >droPer1.super_6 1961352 104 + 6141320 UGACCAGUGUUGCCAACGUUCUUGUU-GUGGCUGACGAUUUGCCAUUUUAAUUAGGCAAUGUCCUGUCAAAAGCAGCCUCUGGCUGUGGCUCUGGUUAUGGGCAG (((((((....(((((((....))))-.((((.((((..(((((..........)))))))))..))))...((((((...))))))))).)))))))....... ( -34.50, z-score = -1.37, R) >droWil1.scaffold_181089 7119866 85 + 12369635 ---------------GUGUUGCUAUU-GCAGUUGACGAUUUGUCAUUUUAAUUAGGCAAUGUCCCGUCAAAGUCAGCCU---CCAGCCAGCAGCAAACUGAAUG- ---------------.(((((((.((-(..((((((((....))..........(((........)))...))))))..---.)))..))))))).........- ( -18.80, z-score = -0.61, R) >consensus CGACGAGUGUUGCCAAAGUCCUUGUUUGUGGCUGACGAUUUGCCAUUUUAAUUAGGCAAUGUCCUGUCACAAGCAGCCG__GCCAGUGGAUACGGUUAUUAGUUA ...........(((...((((.((((..((((.((((..(((((..........)))))))))..))))..))))(((...)))...))))..)))......... (-13.83 = -14.24 + 0.42)
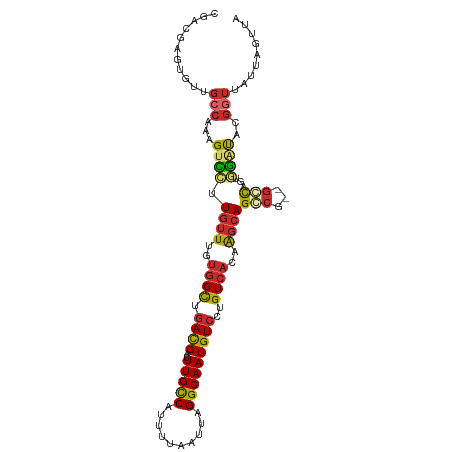


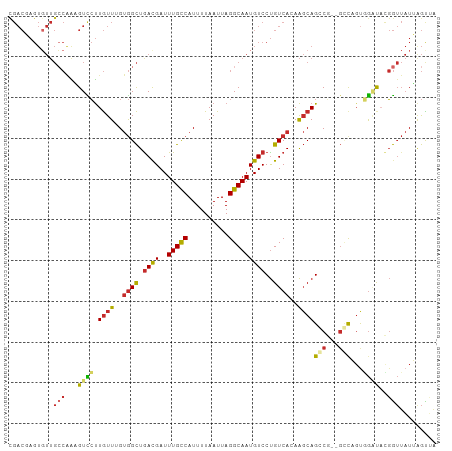
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:56:05 2011