
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 6,580,651 – 6,580,746 |
| Length | 95 |
| Max. P | 0.754048 |

| Location | 6,580,651 – 6,580,746 |
|---|---|
| Length | 95 |
| Sequences | 8 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 73.90 |
| Shannon entropy | 0.47753 |
| G+C content | 0.42579 |
| Mean single sequence MFE | -23.85 |
| Consensus MFE | -13.94 |
| Energy contribution | -13.66 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.34 |
| Structure conservation index | 0.58 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.754048 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
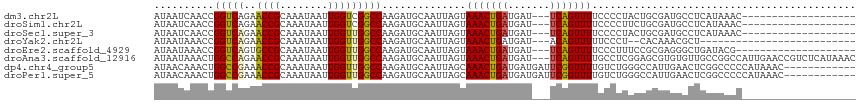
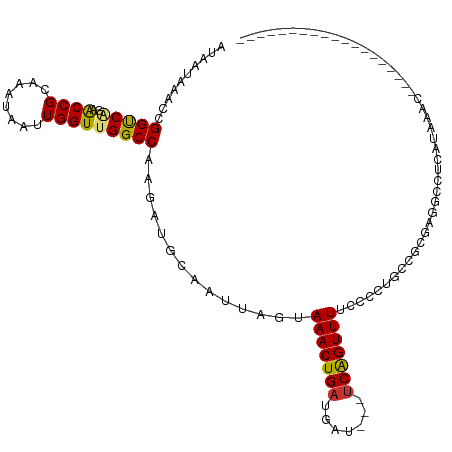
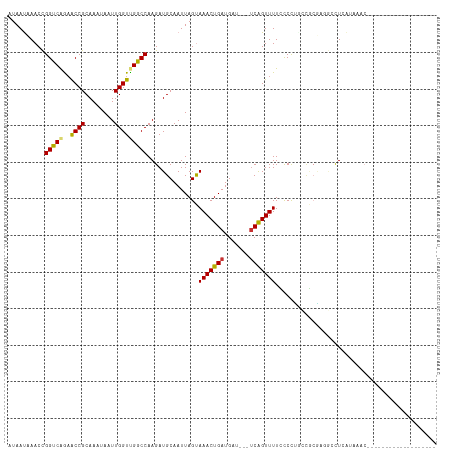
>dm3.chr2L 6580651 95 + 23011544 AUAAUCAACCGGUCAGAACCGCAAAUAAUUGGUCGGCCAAGAUGCAAUUAGUAAACUGAUGAU---UCAGUUUUCCCCUACUGCGAUGCCUCAUAAAC------------------- ..........((((.((.(((........)))))))))..((.(((..(((((((((((....---))))))......)))))...))).))......------------------- ( -19.40, z-score = -1.07, R) >droSim1.chr2L 6373382 95 + 22036055 AUAAUCAACCGGUCAGAACCGCAAAUAAUUGGUCGGCCAAGAUGCAAUUAGUAAACUGAUGAU---UCAGUUUUCCCCUUCUGCGAUGCCUCAUAAAC------------------- ..........((((((((..(((.....((((....))))..))).....(.(((((((....---))))))).)...)))))....)))........------------------- ( -18.60, z-score = -0.70, R) >droSec1.super_3 2120233 95 + 7220098 AUAAUCAACCGGUCAGAACCGCAAAUAAUUGGUUGGCCAAGAUGCAAUUAGUAAACUGAUGAU---UCAGUUUUCCCCUACUGCGAUGCCUCAUAAAC------------------- ..........(((((..((((........)))))))))..((.(((..(((((((((((....---))))))......)))))...))).))......------------------- ( -20.50, z-score = -1.41, R) >droYak2.chr2L 15988908 86 - 22324452 AUAAUAAACCGGUCAGAACCGCAAAUAAUUGGUUGGCCAAGAUGCAAUUAGUAAACUGAUGAU---ACAGUUUUUCCCU--CACAAACGCU-------------------------- ..........((..(((((.(((.....((((....))))..))).(((((....)))))...---...)))))..)).--..........-------------------------- ( -14.00, z-score = -0.16, R) >droEre2.scaffold_4929 15488790 94 + 26641161 AUAAUAAACCGGUCAGUGCCGCAAAUAAUUGGUUGGCCAAGAUGCAAUUAGUAAACUGAUGAU---UCAGUUUUCCCUUUCCGCGAGGGCUGAUACG-------------------- ...........((((((.((((......((((....))))..........(.(((((((....---))))))).).......))).).))))))...-------------------- ( -24.30, z-score = -1.21, R) >droAna3.scaffold_12916 2689152 114 - 16180835 AUAAUAAACUGGCCAGAACCGCAAAUAAUUGGUUGGCCAAGAUGCAAUUAGUAAACUGAUGAU---UCAGUUUUGCCUCGGAGCGUGUGUUGCCGGCCAUUGAACCGUCUCAUAAAC .........(((((..(((((........)))))(((..(.((((..(.((.(((((((....---)))))))...)).)..)))).)...)))))))).................. ( -28.80, z-score = -0.76, R) >dp4.chr4_group5 1180397 105 + 2436548 AUAACAAACUGGCCGAAACCGCAAAUAAUUGGUUGGCCAAGAUGCAAUUAGCAAACUGAUGAUGAUUCGGUUUUGUCUGGGCCAUUGAACUCGGCCCCCAUAAAC------------ ..........(((((((((((........)))))((((.((((((.....))(((((((.......))))))).)))).)))).......)))))).........------------ ( -32.60, z-score = -2.68, R) >droPer1.super_5 5587825 105 + 6813705 AUAACAAACUGGCCGAAACCGCAAAUAAUUGGUUGGCCAAGAUGCAAUUAGCAAACUGAUGAUGAUUCGGUUUUGUCUGGGCCAUUGAACUCGGCCCCCAUAAAC------------ ..........(((((((((((........)))))((((.((((((.....))(((((((.......))))))).)))).)))).......)))))).........------------ ( -32.60, z-score = -2.68, R) >consensus AUAAUAAACCGGUCAGAACCGCAAAUAAUUGGUUGGCCAAGAUGCAAUUAGUAAACUGAUGAU___UCAGUUUUCCCCUGCCGCGAGGCCUCAUAAAC___________________ ..........(((((..((((........)))))))))..............(((((((.......)))))))............................................ (-13.94 = -13.66 + -0.28)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:21:37 2011