
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 3,006,335 – 3,006,447 |
| Length | 112 |
| Max. P | 0.983272 |

| Location | 3,006,335 – 3,006,447 |
|---|---|
| Length | 112 |
| Sequences | 7 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 68.22 |
| Shannon entropy | 0.64277 |
| G+C content | 0.57371 |
| Mean single sequence MFE | -30.59 |
| Consensus MFE | -11.67 |
| Energy contribution | -14.31 |
| Covariance contribution | 2.64 |
| Combinations/Pair | 1.33 |
| Mean z-score | -2.14 |
| Structure conservation index | 0.38 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.88 |
| SVM RNA-class probability | 0.973032 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
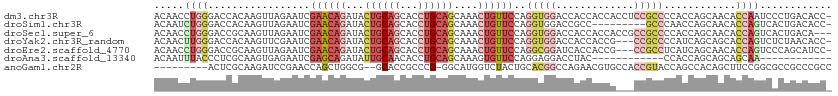
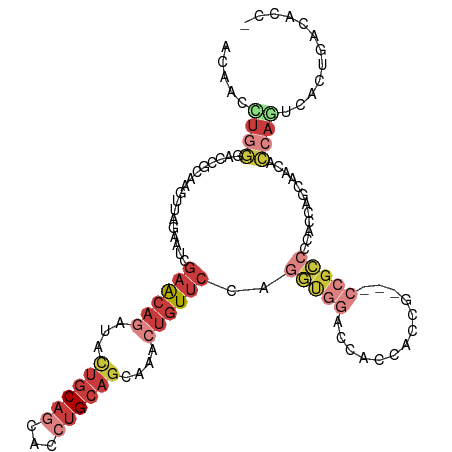
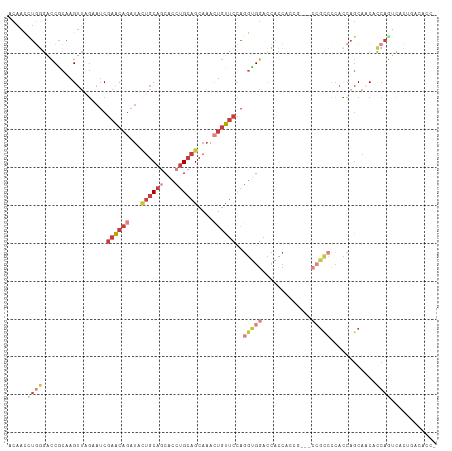
>dm3.chr3R 3006335 112 + 27905053 ACAACCUGGGACCACAAGUUAGAAUCGAACAGAUACUGCAGCACCUGCAGCAAACUGUUCCAGGUGGACCACCACCACCUCCGCCCCACCAGCAACACCAAUCCCUGACACC- .......((((...............((((((...((((((...))))))....))))))..(((((....)))))......((.......))........)))).......- ( -28.50, z-score = -2.59, R) >droSim1.chr3R 3048670 103 + 27517382 ACAAUCUGGGACCACAAGUUAGAAUCGAACAGAUACUGCAGCACCUGCAGCAAACUGUUCCAGGUGGACCGCC---------GCCCAACCAGCAACACCAGUCACUGACACC- .....((((.................((((((...((((((...))))))....))))))..(((((....))---------)))...))))........(((...)))...- ( -27.40, z-score = -1.51, R) >droSec1.super_6 3098555 110 + 4358794 ACAACCUGGGACCGCAAGUUAGAAUCGAACAGAUACUGCAGCACCUGCAGCAAACUGUUCCAGGUGGACCACCACCACCGCCGCCCCACCAGCAACACCAGUCACUGACA--- ......((((.(.((...........((((((...((((((...))))))....))))))..(((((....)))))...)).).)))).(((..((....))..)))...--- ( -33.60, z-score = -2.93, R) >droYak2.chr3R_random 773275 109 + 3277457 ACAACUUGGGACCACAAGUUCGAAUCGAACAGAUACUGCAGCACCUGCAGCAAACUGUUCCAGGUGGACCACCACCG---CCGCCCCAUCAGCAGCACCAGUCUCUAACACC- .......(((((..............((((((...((((((...))))))....))))))..(((((....)))))(---(.((.......)).))....))))).......- ( -31.40, z-score = -2.68, R) >droEre2.scaffold_4770 3268700 109 + 17746568 ACAACCUGGGACCGCAAGUUAGAAUCGAACAGAUACUGCAGCACCUGCAGCAAACUGUUCCAGGCGGAUCACCACCG---CCGCCUCAUCAGCAACACCAGUCCCAGCAUCC- .....((((((((....)...((...((((((...((((((...))))))....))))))..(((((.......)))---))......))..........))))))).....- ( -37.00, z-score = -4.46, R) >droAna3.scaffold_13340 2779710 89 + 23697760 ACAAUUUACCCUCGCAAGUGAGAAUCGAGCAGAUAUUGCAACACCUGCAGCAAAGUGUUCCAGGAGGACCUAC------------CCACCAGCAGCAGCAA------------ ..........((((....)))).(((.....))).((((....((((.(((.....))).)))).((......------------))....))))......------------ ( -16.10, z-score = 0.29, R) >anoGam1.chr2R 23269641 101 + 62725911 ---------ACUCGCAAGAUCCGAACCAGCUGGCG--GCACCGCCCG-GGCAUGGUCUACUGCACGGCCAGAACGUGCCACCGUACCAGCCACAGCUUCCGGCGCCGCCCGCC ---------..((....)).........((.((((--((.(((..(.-(((.(((((....(((((.......)))))....).)))))))...)....))).)))))).)). ( -40.10, z-score = -1.07, R) >consensus ACAACCUGGGACCGCAAGUUAGAAUCGAACAGAUACUGCAGCACCUGCAGCAAACUGUUCCAGGUGGACCACCACCG___CCGCCCCACCAGCAACACCAGUCACUGACACC_ .....((((.................((((((...((((((...))))))....))))))..(((((.............)))))............))))............ (-11.67 = -14.31 + 2.64)

| Location | 3,006,335 – 3,006,447 |
|---|---|
| Length | 112 |
| Sequences | 7 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 68.22 |
| Shannon entropy | 0.64277 |
| G+C content | 0.57371 |
| Mean single sequence MFE | -43.61 |
| Consensus MFE | -18.43 |
| Energy contribution | -20.08 |
| Covariance contribution | 1.64 |
| Combinations/Pair | 1.56 |
| Mean z-score | -2.05 |
| Structure conservation index | 0.42 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.13 |
| SVM RNA-class probability | 0.983272 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
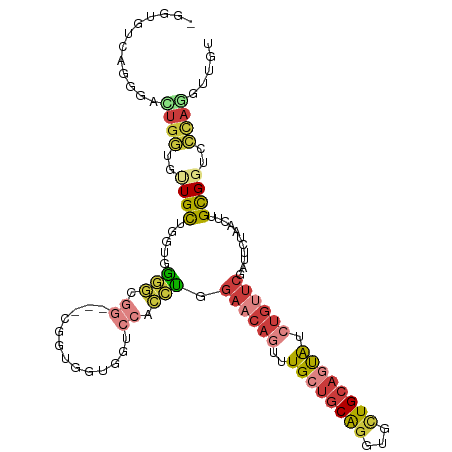
>dm3.chr3R 3006335 112 - 27905053 -GGUGUCAGGGAUUGGUGUUGCUGGUGGGGCGGAGGUGGUGGUGGUCCACCUGGAACAGUUUGCUGCAGGUGCUGCAGUAUCUGUUCGAUUCUAACUUGUGGUCCCAGGUUGU -....((((.(((....))).))))((((((.((((((((((....)))))..((((((..((((((((...)))))))).)))))).......)))).).))))))...... ( -44.40, z-score = -2.23, R) >droSim1.chr3R 3048670 103 - 27517382 -GGUGUCAGUGACUGGUGUUGCUGGUUGGGC---------GGCGGUCCACCUGGAACAGUUUGCUGCAGGUGCUGCAGUAUCUGUUCGAUUCUAACUUGUGGUCCCAGAUUGU -....((((..((....))..))))((((((---------.((((....))..((((((..((((((((...)))))))).))))))...........)).).)))))..... ( -38.60, z-score = -1.40, R) >droSec1.super_6 3098555 110 - 4358794 ---UGUCAGUGACUGGUGUUGCUGGUGGGGCGGCGGUGGUGGUGGUCCACCUGGAACAGUUUGCUGCAGGUGCUGCAGUAUCUGUUCGAUUCUAACUUGCGGUCCCAGGUUGU ---..((((..((....))..))))((((((.((((((((((....)))))..((((((..((((((((...)))))))).)))))).......)).))).))))))...... ( -51.30, z-score = -3.63, R) >droYak2.chr3R_random 773275 109 - 3277457 -GGUGUUAGAGACUGGUGCUGCUGAUGGGGCGG---CGGUGGUGGUCCACCUGGAACAGUUUGCUGCAGGUGCUGCAGUAUCUGUUCGAUUCGAACUUGUGGUCCCAAGUUGU -((..((((...))))..)).....((((((.(---((((((....)))))..((((((..((((((((...)))))))).))))))...........)).))))))...... ( -44.90, z-score = -2.46, R) >droEre2.scaffold_4770 3268700 109 - 17746568 -GGAUGCUGGGACUGGUGUUGCUGAUGAGGCGG---CGGUGGUGAUCCGCCUGGAACAGUUUGCUGCAGGUGCUGCAGUAUCUGUUCGAUUCUAACUUGCGGUCCCAGGUUGU -.....(((((((((..((((((.....)))))---)(((((....)))))..((((((..((((((((...)))))))).))))))............)))))))))..... ( -50.60, z-score = -3.87, R) >droAna3.scaffold_13340 2779710 89 - 23697760 ------------UUGCUGCUGCUGGUGG------------GUAGGUCCUCCUGGAACACUUUGCUGCAGGUGUUGCAAUAUCUGCUCGAUUCUCACUUGCGAGGGUAAAUUGU ------------(((((..(((.(((((------------(..(.(((....))).)...(((..((((((((....)))))))).)))..)))))).)))..)))))..... ( -26.60, z-score = -0.45, R) >anoGam1.chr2R 23269641 101 - 62725911 GGCGGGCGGCGCCGGAAGCUGUGGCUGGUACGGUGGCACGUUCUGGCCGUGCAGUAGACCAUGCC-CGGGCGGUGC--CGCCAGCUGGUUCGGAUCUUGCGAGU--------- (((.(((((((((((..((((((((..(.(((......))).)..)))..)))))...))..((.-...)))))))--)))).)))..((((.......)))).--------- ( -48.90, z-score = -0.28, R) >consensus _GGUGUCAGGGACUGGUGUUGCUGGUGGGGCGG___CGGUGGUGGUCCACCUGGAACAGUUUGCUGCAGGUGCUGCAGUAUCUGUUCGAUUCUAACUUGCGGUCCCAGGUUGU ............((((..((((.....(((.((.............)).))).((((((..((((((((...)))))))).))))))...........))))..))))..... (-18.43 = -20.08 + 1.64)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:55:31 2011