
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 2,890,756 – 2,890,847 |
| Length | 91 |
| Max. P | 0.637441 |

| Location | 2,890,756 – 2,890,847 |
|---|---|
| Length | 91 |
| Sequences | 13 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 59.60 |
| Shannon entropy | 0.88313 |
| G+C content | 0.54860 |
| Mean single sequence MFE | -33.25 |
| Consensus MFE | -5.95 |
| Energy contribution | -6.14 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.70 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.18 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.637441 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 2890756 91 + 27905053 ----------AAAUUGAGUAGGUGGCCGUACGCCAGCCAAUGGAAUAGUGCCCAAGGUGAUUGGCACCUUGCCAGUGAGAUUGCGGUGCGGACCGCUCACC ----------....(((((.((((((.....)))).))...((....(..(((((..(.((((((.....)))))).)..))).))..)...))))))).. ( -33.60, z-score = -1.63, R) >droSim1.chrU 14983550 91 - 15797150 ----------AAAUUGAGUAGGUGGCCGUACGCCAGCCAAUGGAAUAGUGCCCAAGGUGAUUGGCACCUUGCCGGUGAGAUUGCGGUGCGGACCGCUCACC ----------.......((..((((((((((.(((.....)))(((..(((((((((((.....)))))))..))))..)))...)))))).))))..)). ( -34.60, z-score = -1.53, R) >droSec1.super_6 2981099 91 + 4358794 ----------AAAUUGAGUAGGUGGCCGUACGCCAGCCAAUGGAAUAGUGCCCAAGGUGAUUGGCACCUUGCCGGUAAGAUUGCGGUGCGGACCGCUCACC ----------.......((..((((((((((.(((.....)))(((..(((((((((((.....)))))))..))))..)))...)))))).))))..)). ( -35.90, z-score = -2.06, R) >droYak2.chr3R 6663513 89 + 28832112 UAGUUGGCAGGAAGUCUGGUCCGGACACCGCAGUUGCCAGUGGCACGUUGCCAAUGGUCACAGGGAUAAUGAGCCGCGGAUUAGCCAUC------------ ....((((.....(((((...))))).((((.((.((((.((((.....)))).)))).))..((........))))))....))))..------------ ( -33.10, z-score = -1.70, R) >droEre2.scaffold_4770 3144449 91 + 17746568 ----------AAAUUGAGUCGGUGGCUGUACGCCAGCCAAUGGAAUAGUGCCCAAGGUGAUUGGCACCUUGCCGGUGAGAUUGCGGUGCGGACCGCUGACC ----------.......((((((((((((((.(((.....)))(((..(((((((((((.....)))))))..))))..)))...)))))).)))))))). ( -37.60, z-score = -2.57, R) >droAna3.scaffold_13340 2674069 85 + 23697760 ----------------GCCCAGCGGCUUCACGCCCGAUAGUGGAAUAGUGCCUAAUGUGAUAGGCACCUUGCCAGUGAGAUUUCUAUGUGGCCCACUAACA ----------------.....(.(((.....))))(.((((((....(((((((......)))))))...((((...((....))...)))))))))).). ( -28.80, z-score = -2.08, R) >dp4.chr2 11366563 91 - 30794189 ----------AUAUUGAACGGGCGGCCUCAUGCCAGCCAGCGGAAUGGUCCCCAAAGUGAUGGGAAUUUUGCCCGUAAGAUUUCUAUGCGCACCGCUGACU ----------..........((((((.....))).)))(((((..(((...)))..(((((((((((((((....)))))))))))).))).))))).... ( -31.00, z-score = -1.20, R) >droPer1.super_3 5200251 91 - 7375914 ----------AUAUUGAACGGGCGGCCUCAUGCCAGCCAGCGGAAUGGUCCCCAAAGUGAUGGGAAUUUUGCCCGUAAGAUUUCUAUGCGCACCGCUCACU ----------..........((((((.....))).)))(((((..(((...)))..(((((((((((((((....)))))))))))).))).))))).... ( -30.40, z-score = -1.02, R) >droWil1.scaffold_181089 12323363 94 + 12369635 -------CUGUUGCAUUGUUGGUGGCUUAAUACCAGCCAAUGGAAUAGUGCCCAAUGUUAUGGGUAUUUUACCCGACAGAUUACGAUGAGCUCCACUCACU -------(((((((((((((((((......)))))).)))))....((((((((......)))))))).....)))))).......((((.....)))).. ( -28.70, z-score = -2.14, R) >droVir3.scaffold_12855 3627793 89 - 10161210 --------AAACGGUGCGG--CUGGCCCAACGGUAGCCAGGGGCACAUUGCCAAUGGUAAUGGGUAUGACGACCUUGGGAUUGGCCAUCAGCCAGUUCA-- --------....(((.((.--...(((((......((((..(((.....)))..))))..)))))....)))))...((((((((.....)))))))).-- ( -34.40, z-score = -0.80, R) >droMoj3.scaffold_6540 5714868 85 + 34148556 ----------------AACUGGUGGCUUUACGGAAGCCAACGGAAUGGUGCCCAACGUGAUGGGCACUUUGCCAGUGAGAUUACGAUGCGGUCCACUUACA ----------------....(.(((((((...))))))).)((...((((((((......))))))))...))((((.(((..(...)..))))))).... ( -30.10, z-score = -1.90, R) >droGri2.scaffold_14830 2694061 81 - 6267026 --------------------AGUGGCUGUACGGAAGCCAGUGGUAUUGUGCCAAGUGUAAUUGGCACUUUACCGGUGAGAUUACUAUGUGGUCCACUAACA --------------------(.(((((.......))))).)((((..(((((((......)))))))..))))((((.((((((...)))))))))).... ( -33.30, z-score = -3.53, R) >anoGam1.chr2R 50799665 91 - 62725911 ----------GUAGGGUGGCGGUGCCUGGAACGACUCGAGCGGUACCGUACCGAGCACGAUCGGUAUGUUACCGUCCAGGUUGGCAUGCAUGCCGGACGCU ----------...(..(((((.(((.((...((((..(..(((((.((((((((......)))))))).)))))..)..)))).)).))))))))..)... ( -40.70, z-score = -1.71, R) >consensus __________AAAUUGAGUAGGUGGCCGUACGCCAGCCAGUGGAAUAGUGCCCAAGGUGAUGGGCACCUUGCCCGUGAGAUUGCGAUGCGGACCGCUCACC .....................(((......)))........((....(((((..........)))))....))............................ ( -5.95 = -6.14 + 0.19)



| Location | 2,890,756 – 2,890,847 |
|---|---|
| Length | 91 |
| Sequences | 13 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 59.60 |
| Shannon entropy | 0.88313 |
| G+C content | 0.54860 |
| Mean single sequence MFE | -29.72 |
| Consensus MFE | -6.85 |
| Energy contribution | -7.18 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.55 |
| Mean z-score | -1.47 |
| Structure conservation index | 0.23 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.18 |
| SVM RNA-class probability | 0.582620 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 2890756 91 - 27905053 GGUGAGCGGUCCGCACCGCAAUCUCACUGGCAAGGUGCCAAUCACCUUGGGCACUAUUCCAUUGGCUGGCGUACGGCCACCUACUCAAUUU---------- ((((((((((....)))))........((((.....)))).))))).((((......)))).((((((.....))))))............---------- ( -31.60, z-score = -1.51, R) >droSim1.chrU 14983550 91 + 15797150 GGUGAGCGGUCCGCACCGCAAUCUCACCGGCAAGGUGCCAAUCACCUUGGGCACUAUUCCAUUGGCUGGCGUACGGCCACCUACUCAAUUU---------- ((((((((((....)))))....)))))((...((((((..........))))))...))..((((((.....))))))............---------- ( -33.30, z-score = -1.98, R) >droSec1.super_6 2981099 91 - 4358794 GGUGAGCGGUCCGCACCGCAAUCUUACCGGCAAGGUGCCAAUCACCUUGGGCACUAUUCCAUUGGCUGGCGUACGGCCACCUACUCAAUUU---------- ((((((((((....)))))....)))))((...((((((..........))))))...))..((((((.....))))))............---------- ( -31.10, z-score = -1.28, R) >droYak2.chr3R 6663513 89 - 28832112 ------------GAUGGCUAAUCCGCGGCUCAUUAUCCCUGUGACCAUUGGCAACGUGCCACUGGCAACUGCGGUGUCCGGACCAGACUUCCUGCCAACUA ------------..((((....((((((.((((.......))))(((.((((.....)))).)))...)))))).....(((.......))).)))).... ( -27.60, z-score = -1.01, R) >droEre2.scaffold_4770 3144449 91 - 17746568 GGUCAGCGGUCCGCACCGCAAUCUCACCGGCAAGGUGCCAAUCACCUUGGGCACUAUUCCAUUGGCUGGCGUACAGCCACCGACUCAAUUU---------- ((((.(((((....))))).........(((((((((.....))))))).............((((((.....))))))))))))......---------- ( -32.10, z-score = -2.23, R) >droAna3.scaffold_13340 2674069 85 - 23697760 UGUUAGUGGGCCACAUAGAAAUCUCACUGGCAAGGUGCCUAUCACAUUAGGCACUAUUCCACUAUCGGGCGUGAAGCCGCUGGGC---------------- ...((((((((((...((....))...))))..((((((((......))))))))...))))))((.((((......)))).)).---------------- ( -32.80, z-score = -2.79, R) >dp4.chr2 11366563 91 + 30794189 AGUCAGCGGUGCGCAUAGAAAUCUUACGGGCAAAAUUCCCAUCACUUUGGGGACCAUUCCGCUGGCUGGCAUGAGGCCGCCCGUUCAAUAU---------- ((((((((((((.(..((....))...).)))...((((((......)))))).....)))))))))(((.....))).............---------- ( -31.40, z-score = -1.24, R) >droPer1.super_3 5200251 91 + 7375914 AGUGAGCGGUGCGCAUAGAAAUCUUACGGGCAAAAUUCCCAUCACUUUGGGGACCAUUCCGCUGGCUGGCAUGAGGCCGCCCGUUCAAUAU---------- ..(((((((.((..............((((.....((((((......))))))....))))..((((.......)))))))))))))....---------- ( -30.50, z-score = -0.72, R) >droWil1.scaffold_181089 12323363 94 - 12369635 AGUGAGUGGAGCUCAUCGUAAUCUGUCGGGUAAAAUACCCAUAACAUUGGGCACUAUUCCAUUGGCUGGUAUUAAGCCACCAACAAUGCAACAG------- .(.((((((.(((((..((....(((.(((((...))))))))))..))))).))))))).((((.((((.....))))))))...........------- ( -26.30, z-score = -1.52, R) >droVir3.scaffold_12855 3627793 89 + 10161210 --UGAACUGGCUGAUGGCCAAUCCCAAGGUCGUCAUACCCAUUACCAUUGGCAAUGUGCCCCUGGCUACCGUUGGGCCAG--CCGCACCGUUU-------- --.(((((((.((((((((........))))))))...)))..............((((..((((((.......))))))--..)))).))))-------- ( -28.60, z-score = -0.24, R) >droMoj3.scaffold_6540 5714868 85 - 34148556 UGUAAGUGGACCGCAUCGUAAUCUCACUGGCAAAGUGCCCAUCACGUUGGGCACCAUUCCGUUGGCUUCCGUAAAGCCACCAGUU---------------- (((.((((((..((...))..)).)))).)))..(((((((......)))))))......(.((((((.....)))))).)....---------------- ( -26.20, z-score = -1.68, R) >droGri2.scaffold_14830 2694061 81 + 6267026 UGUUAGUGGACCACAUAGUAAUCUCACCGGUAAAGUGCCAAUUACACUUGGCACAAUACCACUGGCUUCCGUACAGCCACU-------------------- .(((((((((((....((....))....)))...(((((((......)))))))....))))))))...............-------------------- ( -23.80, z-score = -2.40, R) >anoGam1.chr2R 50799665 91 + 62725911 AGCGUCCGGCAUGCAUGCCAACCUGGACGGUAACAUACCGAUCGUGCUCGGUACGGUACCGCUCGAGUCGUUCCAGGCACCGCCACCCUAC---------- .(((...((((....))))..(((((((((((...)))))..((.((((((..((....)).)))))))).))))))...)))........---------- ( -31.00, z-score = -0.49, R) >consensus AGUGAGCGGUCCGCAUCGCAAUCUCACCGGCAAAGUGCCAAUCACCUUGGGCACUAUUCCACUGGCUGGCGUACAGCCACCAACUCAAUUU__________ ............................((.....((((..........)))).....))..(((((.......)))))...................... ( -6.85 = -7.18 + 0.33)

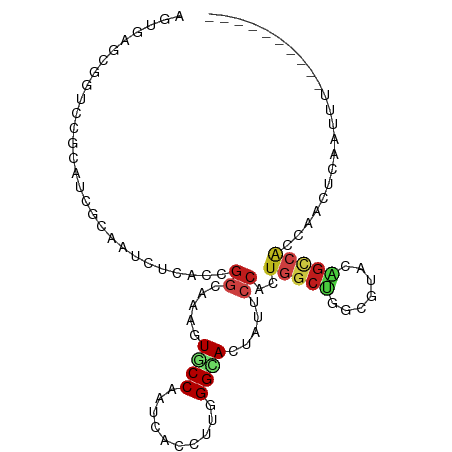

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:55:19 2011