
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 6,516,252 – 6,516,358 |
| Length | 106 |
| Max. P | 0.803105 |

| Location | 6,516,252 – 6,516,350 |
|---|---|
| Length | 98 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 69.32 |
| Shannon entropy | 0.54465 |
| G+C content | 0.54455 |
| Mean single sequence MFE | -33.77 |
| Consensus MFE | -15.31 |
| Energy contribution | -16.70 |
| Covariance contribution | 1.39 |
| Combinations/Pair | 1.30 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.74 |
| SVM RNA-class probability | 0.803105 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
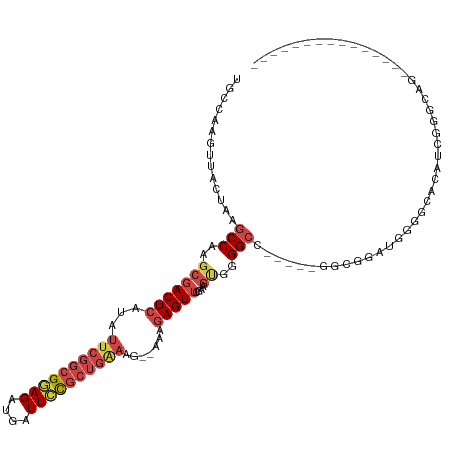
>dm3.chr2L 6516252 98 - 23011544 UGCCAAGUUACUAAGCCAAGCGACUCAUAUUCGGCGAAGAUGAUUUCGCUGAAAG--AAAGAGUUCGAAGUGGGGCC-----GACGGAUGGGGCACAUCGGGCAG--------------- ((((..........(((..(((((((...((((((((((....))))))))))..--...)))))....))..))).-----....((((.....)))).)))).--------------- ( -31.10, z-score = -1.57, R) >droSim1.chr2L 6307593 100 - 22036055 UGCCAAGUUACUAAGCCAAGCGACUCAUAUUCGGCGGAGAUGAUUCCGCUGAAAGAAAAAGAGUUCGAAGUGGGGCC-----UGCGGGUGGGGCACAUCGGGCAG--------------- ((((...........(((..((((((...((((((((((....)))))))))).......))).)))...))).(((-----(.......))))......)))).--------------- ( -32.80, z-score = -1.02, R) >droSec1.super_3 2056692 100 - 7220098 UGCCAAGUUACUAAGCCAAGCGACUCAUAUUCGGCGGAGGUGAUUCCGCUGAAAGAAAAAGAGUUCGAAGUGGGGCC-----GGCGGAUGGGGCACAUCGGGCAG--------------- ((((..........(((..((..(((((.((((((((((....)))))))))).(((......)))...))))))).-----))).((((.....)))).)))).--------------- ( -35.10, z-score = -1.79, R) >droYak2.chr2L 15924537 103 + 22324452 UGCCAAGUUACUAAGCCAAGCGACUCGUAUUCGGCGGAGAUGAUUCCGCUGAAAG--AAAGAGUUCAAAGCGGGGCUCAUCGGGCGGUUGGGGUACAUGGGGCAG--------------- ((((..((((((...((((.((.((((..((((((((((....))))))))))..--...((((((......))))))..)))))).)))))))).))..)))).--------------- ( -41.50, z-score = -3.58, R) >droEre2.scaffold_4929 15424681 85 - 26641161 UGCCAAGUUACUAAGCCAAGCGACUCAUAUUCGGCGGAGAUGAUUCCGCUGGAAG--AAAGAGUUCGAAGUGGGGCCCAUCGGGCAG--------------------------------- ...............(((..((((((...((((((((((....))))))))))..--...))).)))...))).((((...))))..--------------------------------- ( -28.40, z-score = -1.49, R) >droAna3.scaffold_12916 2620442 109 + 16180835 ---CAGGCCUUCAGUCCCAUCCACUAUUA------GUAGAUGCUUUUGCUGAAAG--AAAGAGUUGCAACUAGGGCGGAUGGGAUGGAUAGGGUGGUGUGUGGGCUGUGGUGGGCAGAGG ---...((((.((((((((((((((.(((------(((((....))))))))...--.....(((.((.((.....)).)).)))......))))).))).))))))....))))..... ( -33.70, z-score = -0.35, R) >consensus UGCCAAGUUACUAAGCCAAGCGACUCAUAUUCGGCGGAGAUGAUUCCGCUGAAAG__AAAGAGUUCGAAGUGGGGCC_____GGCGGAUGGGGCACAUCGGGCAG_______________ ......(((.....(((..(((((((...((((((((((....)))))))))).......)))))....))..)))......)))................................... (-15.31 = -16.70 + 1.39)



| Location | 6,516,262 – 6,516,358 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 72.12 |
| Shannon entropy | 0.49402 |
| G+C content | 0.54049 |
| Mean single sequence MFE | -33.02 |
| Consensus MFE | -17.80 |
| Energy contribution | -19.55 |
| Covariance contribution | 1.75 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.14 |
| Structure conservation index | 0.54 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.04 |
| SVM RNA-class probability | 0.513563 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 6516262 96 - 23011544 GGCCAUGUUGCCAAGUUACUAAGCCAAGCGACUCAUAUUCGGCGAAGAUGAUUUCGCUGAAAG--AAAGAGUUCGAAGUGGGGCCGACGGAUGGGGCA--------------- .(((.(((((((.((...))...(((..((((((...((((((((((....))))))))))..--...))).)))...))))).))))).....))).--------------- ( -29.60, z-score = -0.89, R) >droSim1.chr2L 6307603 98 - 22036055 GGCCAUGUUGCCAAGUUACUAAGCCAAGCGACUCAUAUUCGGCGGAGAUGAUUCCGCUGAAAGAAAAAGAGUUCGAAGUGGGGCCUGCGGGUGGGGCA--------------- (((......)))..........(((......(((((.((((((((((....)))))))))).(((......)))...)))))(((....)))..))).--------------- ( -30.70, z-score = -0.16, R) >droSec1.super_3 2056702 98 - 7220098 GGCCAUGUUGCCAAGUUACUAAGCCAAGCGACUCAUAUUCGGCGGAGGUGAUUCCGCUGAAAGAAAAAGAGUUCGAAGUGGGGCCGGCGGAUGGGGCA--------------- .(((...(((((.((...))..(((..(((((((...((((((((((....)))))))))).......)))))....))..))).)))))....))).--------------- ( -33.20, z-score = -0.86, R) >droYak2.chr2L 15924547 101 + 22324452 GGCCAUGUUGCCAAGUUACUAAGCCAAGCGACUCGUAUUCGGCGGAGAUGAUUCCGCUGAAAG--AAAGAGUUCAAAGCGGGGCUCAUCGGGCGGUUGGGGUA---------- (((......)))....((((...((((.((.((((..((((((((((....))))))))))..--...((((((......))))))..)))))).))))))))---------- ( -37.40, z-score = -1.88, R) >droEre2.scaffold_4929 15424683 91 - 26641161 GGCCGUGUUGCCAAGUUACUAAGCCAAGCGACUCAUAUUCGGCGGAGAUGAUUCCGCUGGAAG--AAAGAGUUCGAAGUGGGGCCCAUCGGGC-------------------- (((......)))...........(((..((((((...((((((((((....))))))))))..--...))).)))...))).((((...))))-------------------- ( -30.50, z-score = -0.89, R) >droAna3.scaffold_12916 2620457 99 + 16180835 -----GGCUG-CAGGCCUUCAGUCCCAUCCACUAUUAGU------AGAUGCUUUUGCUGAAAG--AAAGAGUUGCAACUAGGGCGGAUGGGAUGGAUAGGGUGGUGUGUGGGC -----(.(..-((.(((.((.(((((((((.((.(((((------(((....)))))))).))--........((.......))))))))))).))......))).))..).) ( -36.70, z-score = -2.15, R) >consensus GGCCAUGUUGCCAAGUUACUAAGCCAAGCGACUCAUAUUCGGCGGAGAUGAUUCCGCUGAAAG__AAAGAGUUCGAAGUGGGGCCGACGGGUGGGGCA_______________ ((((..(((((...(((....)))...))))).....((((((((((....))))))))))....................))))............................ (-17.80 = -19.55 + 1.75)


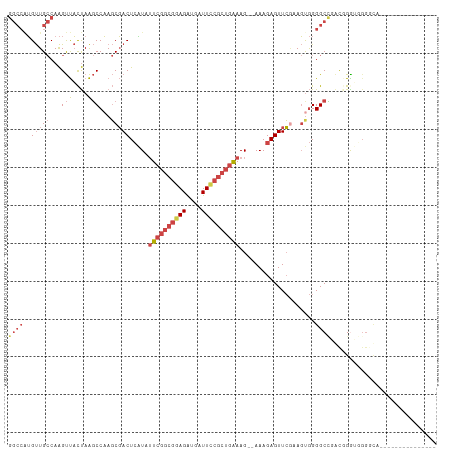
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:21:33 2011