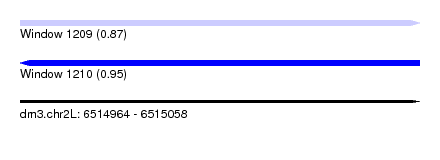

| Sequence ID | dm3.chr2L |
|---|---|
| Location | 6,514,964 – 6,515,058 |
| Length | 94 |
| Max. P | 0.954324 |

| Location | 6,514,964 – 6,515,058 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 67.10 |
| Shannon entropy | 0.60595 |
| G+C content | 0.54820 |
| Mean single sequence MFE | -27.07 |
| Consensus MFE | -17.08 |
| Energy contribution | -17.83 |
| Covariance contribution | 0.76 |
| Combinations/Pair | 1.48 |
| Mean z-score | -0.75 |
| Structure conservation index | 0.63 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.00 |
| SVM RNA-class probability | 0.872064 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
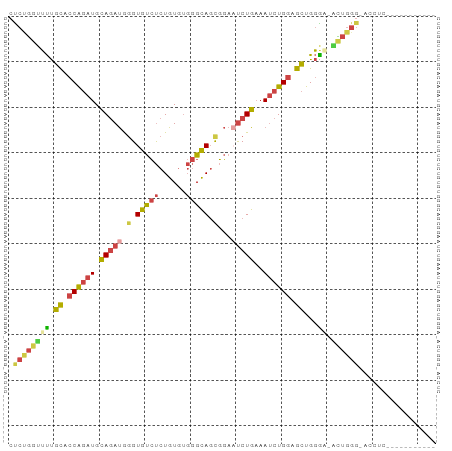
>dm3.chr2L 6514964 94 + 23011544 CUCUGGUUUUGCACCAGAUGCAGAUGGGUGUCUCUGUGUGGGCAGCGGAAUCUGAAAUCUGGAGCUGGAAUACUGGGGAGCUUGGCUUGGCUGG (((..((...((.((((((.(((((.(.(((((......))))).)...)))))..)))))).))......))..)))((((......)))).. ( -32.20, z-score = -1.15, R) >droSim1.chr2L 6306292 82 + 22036055 CUCUGGUUUUGCACCAGAUGCAGAUGGGUGUCUCUGUGUGGGCAGUGGAAUCUGAAAUCUGGAGCUGGGC-ACUGGGUACUGC----------- (((..(((..((.((((((.(((((.(.(((((......))))).)...)))))..)))))).))..)))-...)))......----------- ( -25.90, z-score = -0.41, R) >droSec1.super_3 2055389 82 + 7220098 CUCUGGUUUUGCACCAGAUGCAGAUGGGUGUCUCUGUGUGGGCAGUGGAAUCUGAAAUCUGGAGCUGGGC-ACUGGGUACUGC----------- (((..(((..((.((((((.(((((.(.(((((......))))).)...)))))..)))))).))..)))-...)))......----------- ( -25.90, z-score = -0.41, R) >droYak2.chr2L 15923252 81 - 22324452 CUCUGGUUUUGCACCAGAUGCAGAUGGGUGUCUCUGUGUGGGCAGCGGAUUCUGAAAUCUGGAGCUGGAA-GCUGGG-AGCUU----------- .((..(((((((.((((((.(((((.(.(((((......))))).).)..))))..)))))).))..)))-))..))-.....----------- ( -27.20, z-score = -0.63, R) >droEre2.scaffold_4929 15423397 74 + 26641161 CUCUGGGUUUGCGCCAGAUGCAGAUGGGUGUCUCUGUGUGGGCAGUGGAAUCUGAAAUCUGGAGCUGGGAGCUU-------------------- ((((.((((....((((((.(((((.(.(((((......))))).)...)))))..)))))))))).))))...-------------------- ( -24.90, z-score = -1.10, R) >dp4.chr4_group1 1576506 85 + 5278887 GCCCCGUGCCUUCUCGCAUGUACACAAGUACACAUGUA---GUACCUUCUCCUGCCACAUGCAGAUGUGUCUGUGGGGCACUCUGUGU------ (((((((((......)))))..((((..(((((((((.---(((........))).)))))....))))..)))))))).........------ ( -26.30, z-score = -0.78, R) >consensus CUCUGGUUUUGCACCAGAUGCAGAUGGGUGUCUCUGUGUGGGCAGCGGAAUCUGAAAUCUGGAGCUGGGA_ACUGGG_ACCUC___________ .((((((...((.((((((.(((((.(.(((((......))))).)...)))))..)))))).))......))))))................. (-17.08 = -17.83 + 0.76)
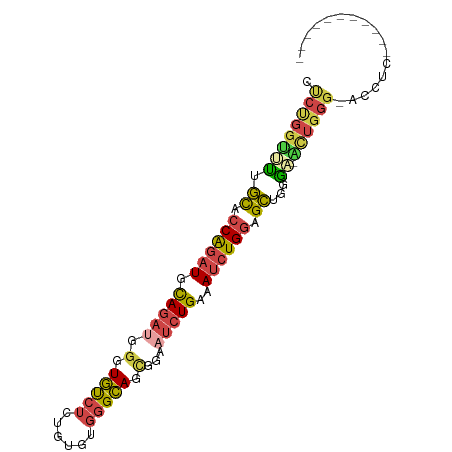


| Location | 6,514,964 – 6,515,058 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 67.10 |
| Shannon entropy | 0.60595 |
| G+C content | 0.54820 |
| Mean single sequence MFE | -19.18 |
| Consensus MFE | -10.80 |
| Energy contribution | -10.75 |
| Covariance contribution | -0.05 |
| Combinations/Pair | 1.35 |
| Mean z-score | -1.30 |
| Structure conservation index | 0.56 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.60 |
| SVM RNA-class probability | 0.954324 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
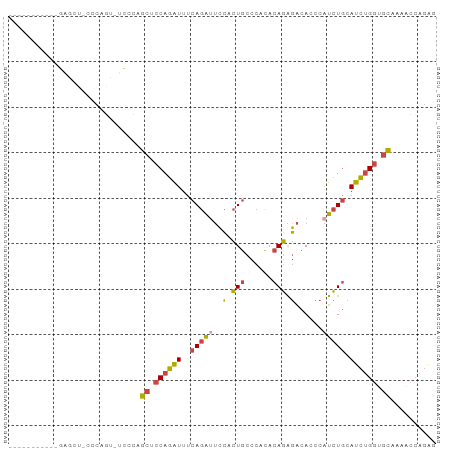
>dm3.chr2L 6514964 94 - 23011544 CCAGCCAAGCCAAGCUCCCCAGUAUUCCAGCUCCAGAUUUCAGAUUCCGCUGCCCACACAGAGACACCCAUCUGCAUCUGGUGCAAAACCAGAG ..(((........))).............((.((((((..(((((((..(((......))).)).....))))).)))))).)).......... ( -17.80, z-score = -0.99, R) >droSim1.chr2L 6306292 82 - 22036055 -----------GCAGUACCCAGU-GCCCAGCUCCAGAUUUCAGAUUCCACUGCCCACACAGAGACACCCAUCUGCAUCUGGUGCAAAACCAGAG -----------(((.(....).)-))...((.((((((..(((((((..(((......))).)).....))))).)))))).)).......... ( -17.90, z-score = -1.40, R) >droSec1.super_3 2055389 82 - 7220098 -----------GCAGUACCCAGU-GCCCAGCUCCAGAUUUCAGAUUCCACUGCCCACACAGAGACACCCAUCUGCAUCUGGUGCAAAACCAGAG -----------(((.(....).)-))...((.((((((..(((((((..(((......))).)).....))))).)))))).)).......... ( -17.90, z-score = -1.40, R) >droYak2.chr2L 15923252 81 + 22324452 -----------AAGCU-CCCAGC-UUCCAGCUCCAGAUUUCAGAAUCCGCUGCCCACACAGAGACACCCAUCUGCAUCUGGUGCAAAACCAGAG -----------((((.-....))-)).((((....((((....)))).))))......((((........))))..((((((.....)))))). ( -17.90, z-score = -1.26, R) >droEre2.scaffold_4929 15423397 74 - 26641161 --------------------AAGCUCCCAGCUCCAGAUUUCAGAUUCCACUGCCCACACAGAGACACCCAUCUGCAUCUGGCGCAAACCCAGAG --------------------.........((.((((((..(((((((..(((......))).)).....))))).)))))).)).......... ( -16.20, z-score = -2.09, R) >dp4.chr4_group1 1576506 85 - 5278887 ------ACACAGAGUGCCCCACAGACACAUCUGCAUGUGGCAGGAGAAGGUAC---UACAUGUGUACUUGUGUACAUGCGAGAAGGCACGGGGC ------.........(((((....(((((..((((((((..((.........)---).))))))))..)))))...(((......))).))))) ( -27.40, z-score = -0.67, R) >consensus ___________GAGCU_CCCAGU_UCCCAGCUCCAGAUUUCAGAUUCCACUGCCCACACAGAGACACCCAUCUGCAUCUGGUGCAAAACCAGAG .............................((.((((((..(((((..(.(((......))).)......))))).)))))).)).......... (-10.80 = -10.75 + -0.05)
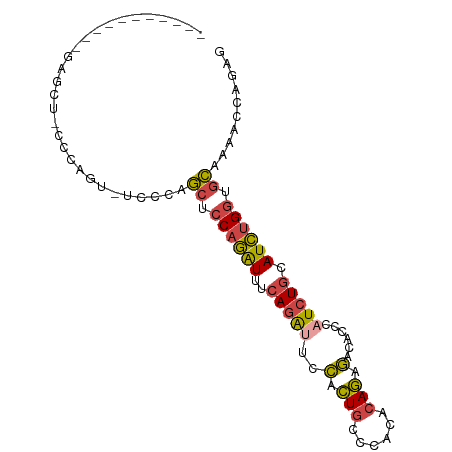


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:21:31 2011