
| Sequence ID | dm3.chr2L |
|---|---|
| Location | 6,511,119 – 6,511,222 |
| Length | 103 |
| Max. P | 0.805654 |

| Location | 6,511,119 – 6,511,222 |
|---|---|
| Length | 103 |
| Sequences | 8 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 83.20 |
| Shannon entropy | 0.32655 |
| G+C content | 0.51066 |
| Mean single sequence MFE | -28.79 |
| Consensus MFE | -22.97 |
| Energy contribution | -23.16 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.41 |
| Structure conservation index | 0.80 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.75 |
| SVM RNA-class probability | 0.805654 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr2L 6511119 103 - 23011544 -----CUACGCCACUGACACGUACUAACGAGCGACGCCUGACAGCUCGCAGAGAGCGUCA-UCGUCCCGCUCGACGAUGCUUAAAACUUAAUUUGAGGCAACAAAGACA------- -----....(((........(((....(((((((((..((((.((((.....))))))))-.))))..)))))....)))(((((......))))))))..........------- ( -28.70, z-score = -1.75, R) >droSim1.chr2L 6302389 103 - 22036055 -----CUACGCCACUGACACGUACUAACGAGCGACGCCUGACAGCUCGCAGAGAGCGUCA-UCGUCCCGCUCGACGAUGCUUAAAACUUAAUUUGAGGCAACAAAGACA------- -----....(((........(((....(((((((((..((((.((((.....))))))))-.))))..)))))....)))(((((......))))))))..........------- ( -28.70, z-score = -1.75, R) >droSec1.super_3 2051713 103 - 7220098 -----CUACGCCACUAACACGUACUAACGAGCGACGCCUGACAGCUCGCAGAGAGCGUCA-UCGUCCCGCUCGACGAUGCUUAAAACUUAGUUUGAGGCAACAAAGACA------- -----....(((...............(((((((((..((((.((((.....))))))))-.))))..))))).(((.(((........)))))).)))..........------- ( -29.80, z-score = -1.82, R) >droYak2.chr2L 15919409 104 + 22324452 -----------CACUGACACGUACUAACGAGCAACGCCUGACAGCUCGCAGAGAGCGUCA-UCGUCCCGCUCGACGAUGCUUAAAACUUAAUUUGAGGCAACAAAGACGUAGGUCU -----------....(((((((.((..(((((.(((..((((.((((.....))))))))-.)))...)))))..(.(((((.(((.....))).))))).)..))))))..))). ( -29.50, z-score = -1.34, R) >droEre2.scaffold_4929 15419580 104 - 26641161 -----------CACUGACACGUACUAACGAGCGACGCCUGACAGCUCGUAGAGAGCGUCA-UCGUCCCGCUCGACGAUGCUUAAAACUUAAUUUGAGGCAACAAAGACUGAGCUCU -----------.................((((...(((((((.((((.....)))))))(-(((((......)))))).................))))..((.....)).)))). ( -28.00, z-score = -0.93, R) >droAna3.scaffold_12916 2614676 115 + 16180835 AUAUUGUGCGCCACUGACACGUACUAACGAGCGCCGCUUGACAGCUCGCAG-GAGCGUCGUUCGCCCCGCCUGGAGAUGCUUAAAACUUAAUUUGAAGCAACAAAGACAAGGAUCU .....(.((((.((......)).(....).))))).((((....(((.(((-(.(((.....)))....))))))).(((((.(((.....))).))))).......))))..... ( -24.50, z-score = 1.62, R) >dp4.chr4_group1 1571404 101 - 5278887 ----AUAGUGCGACUGACACGUACUAACGAGCGACGCCUGACAGC---GAAAGAGCGUCA-UCGUCCCGCUCGACGAUGCUUAAAACUUAAUUUGAAGCAACAAAAGAC------- ----.(((((((.......))))))).(((((((((..((((.((---......))))))-.))))..)))))..(.(((((.(((.....))).))))).).......------- ( -29.90, z-score = -2.80, R) >droPer1.super_5 3178566 101 + 6813705 ----AUAGUGCGGCUGACACGUACUAACGAGCGACGCCUGACAGC---GAAAGAGCGUCA-UCGUCCCGCUCGACGAUGCUUAAAACUUAAUUUGAAGCAACAAAAGAC------- ----.(((((((.......))))))).(((((((((..((((.((---......))))))-.))))..)))))..(.(((((.(((.....))).))))).).......------- ( -31.20, z-score = -2.54, R) >consensus _____CUACGCCACUGACACGUACUAACGAGCGACGCCUGACAGCUCGCAGAGAGCGUCA_UCGUCCCGCUCGACGAUGCUUAAAACUUAAUUUGAGGCAACAAAGACA_______ ...........................((((((..((.((((.((((.....))))))))...))..))))))..(.(((((.(((.....))).))))).).............. (-22.97 = -23.16 + 0.19)
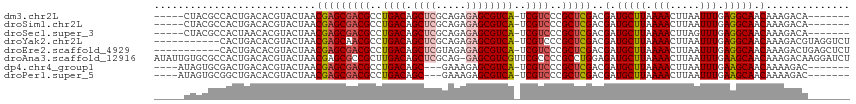


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:21:30 2011