
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 2,736,041 – 2,736,138 |
| Length | 97 |
| Max. P | 0.958148 |

| Location | 2,736,041 – 2,736,138 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 88.87 |
| Shannon entropy | 0.19971 |
| G+C content | 0.44178 |
| Mean single sequence MFE | -18.28 |
| Consensus MFE | -13.51 |
| Energy contribution | -15.20 |
| Covariance contribution | 1.69 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.47 |
| Structure conservation index | 0.74 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.29 |
| SVM RNA-class probability | 0.921541 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
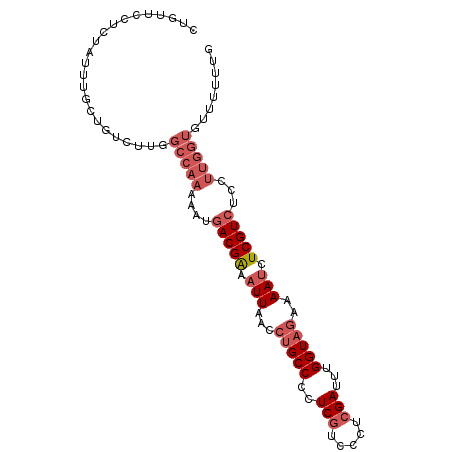
>dm3.chr3R 2736041 97 + 27905053 CUGUUCCUCUAUUUGCUGUCUUGGCCAAAAAUGACGAAAUUAACCUGCCCCUCGUCUCUCGAUUUGGUAGAAAAUCUCGUCUCCUUGGUGUUUUUUG .......................(((((....(((((.(((...(((((..(((.....)))...)))))..))).)))))...)))))........ ( -20.00, z-score = -3.01, R) >droEre2.scaffold_4770 2987270 87 + 17746568 CUGUUCCUCUAUUUGCUGUCUUG----------ACGAAAUUAUCCUGCCCCUCGUCCCUCGAUUUGGUAGAAAAUCUCGUCUCCUUGGUGUUCUUUG ..............((((....(----------((((.(((...(((((..(((.....)))...)))))..))).)))))....))))........ ( -16.40, z-score = -3.29, R) >droYak2.chr3R 21156809 92 - 28832112 CUGUUCCUCUAUUUGCUGUCUUGGCCAAAAAUGACGAAAUUA----GCCCCUCGUCUCUCGAUUUGGUA-AAAACCUCGUCUCCUUGGUGUUCUUUG .......................(((((....(((((.....----(((..(((.....)))...))).-......)))))...)))))........ ( -14.50, z-score = -1.22, R) >droSec1.super_6 2827571 96 + 4358794 CUGUUCCUCUAUUUGCUGUCUUGGCCAAAUAUGACGGAAUUAACCUGCCACUCCUCCCUCGAUUUGGUAGAAAAUCUCGUCUACUUGGUGUUUUUG- .......................(((((.((.((((..(((...((((((.((.......))..))))))..)))..)))))).))))).......- ( -19.10, z-score = -1.74, R) >droSim1.chr3R 2772759 97 + 27517382 CUGUUCCUCUAUUUGCUGUCUUGGCCAAAAAUGACGAAAUUAACCUGCCACUCGUCCCUCGAUUUGGUAGAAAAUCUCGUCUCCUUGGUGUUUUUUG .......................(((((....(((((.(((...((((((.(((.....)))..))))))..))).)))))...)))))........ ( -21.40, z-score = -3.10, R) >consensus CUGUUCCUCUAUUUGCUGUCUUGGCCAAAAAUGACGAAAUUAACCUGCCCCUCGUCCCUCGAUUUGGUAGAAAAUCUCGUCUCCUUGGUGUUUUUUG .......................(((((....(((((.(((...(((((..(((.....)))...)))))..))).)))))...)))))........ (-13.51 = -15.20 + 1.69)


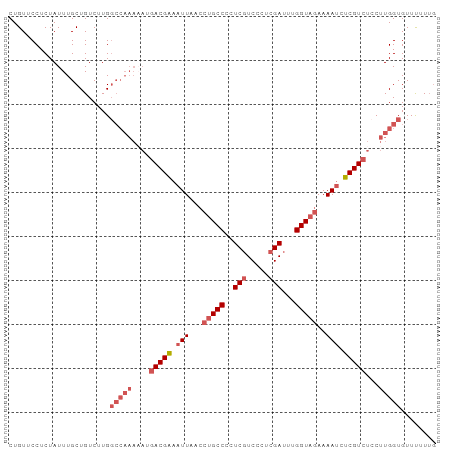
| Location | 2,736,041 – 2,736,138 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 88.87 |
| Shannon entropy | 0.19971 |
| G+C content | 0.44178 |
| Mean single sequence MFE | -21.70 |
| Consensus MFE | -15.67 |
| Energy contribution | -17.52 |
| Covariance contribution | 1.85 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.77 |
| Structure conservation index | 0.72 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.65 |
| SVM RNA-class probability | 0.958148 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
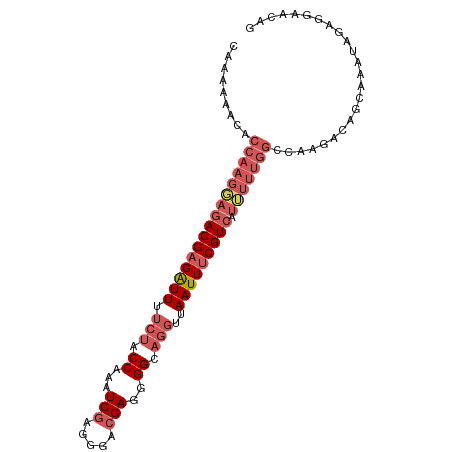
>dm3.chr3R 2736041 97 - 27905053 CAAAAAACACCAAGGAGACGAGAUUUUCUACCAAAUCGAGAGACGAGGGGCAGGUUAAUUUCGUCAUUUUUGGCCAAGACAGCAAAUAGAGGAACAG .........((((((((((((((((.(((.((...(((.....)))..)).)))..))))))))).)))))))........................ ( -23.80, z-score = -3.40, R) >droEre2.scaffold_4770 2987270 87 - 17746568 CAAAGAACACCAAGGAGACGAGAUUUUCUACCAAAUCGAGGGACGAGGGGCAGGAUAAUUUCGU----------CAAGACAGCAAAUAGAGGAACAG ................(((((((((((((.((...(((.....)))..)).)))).))))))))----------)...................... ( -18.90, z-score = -3.13, R) >droYak2.chr3R 21156809 92 + 28832112 CAAAGAACACCAAGGAGACGAGGUUUU-UACCAAAUCGAGAGACGAGGGGC----UAAUUUCGUCAUUUUUGGCCAAGACAGCAAAUAGAGGAACAG .........((((((((((((((((..-..((...(((.....))).))..----.))))))))).)))))))........................ ( -21.50, z-score = -2.76, R) >droSec1.super_6 2827571 96 - 4358794 -CAAAAACACCAAGUAGACGAGAUUUUCUACCAAAUCGAGGGAGGAGUGGCAGGUUAAUUCCGUCAUAUUUGGCCAAGACAGCAAAUAGAGGAACAG -........(((((((((((..(((.(((.(((..((......))..))).)))..)))..)))).)))))))........................ ( -20.80, z-score = -1.81, R) >droSim1.chr3R 2772759 97 - 27517382 CAAAAAACACCAAGGAGACGAGAUUUUCUACCAAAUCGAGGGACGAGUGGCAGGUUAAUUUCGUCAUUUUUGGCCAAGACAGCAAAUAGAGGAACAG .........((((((((((((((((.(((.(((..(((.....))).))).)))..))))))))).)))))))........................ ( -23.50, z-score = -2.75, R) >consensus CAAAAAACACCAAGGAGACGAGAUUUUCUACCAAAUCGAGGGACGAGGGGCAGGUUAAUUUCGUCAUUUUUGGCCAAGACAGCAAAUAGAGGAACAG .........((((((((((((((((..((.((...(((.....)))..)).))...))))))))).)))))))........................ (-15.67 = -17.52 + 1.85)

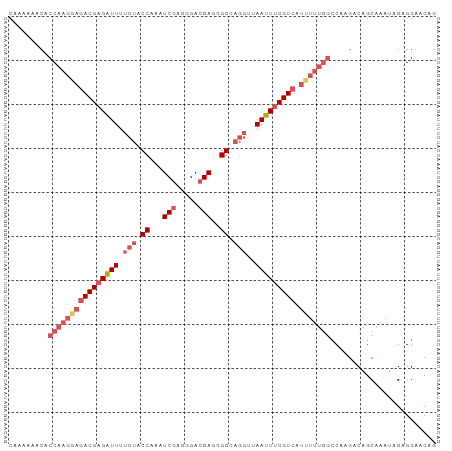
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:54:55 2011