| Sequence ID | dm3.chr3R |
|---|---|
| Location | 2,658,187 – 2,658,277 |
| Length | 90 |
| Max. P | 0.991209 |

| Location | 2,658,187 – 2,658,277 |
|---|---|
| Length | 90 |
| Sequences | 9 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 86.25 |
| Shannon entropy | 0.25322 |
| G+C content | 0.52572 |
| Mean single sequence MFE | -35.79 |
| Consensus MFE | -29.93 |
| Energy contribution | -29.93 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.67 |
| Structure conservation index | 0.84 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.46 |
| SVM RNA-class probability | 0.991209 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 2658187 90 + 27905053 ----GGGGAGUGUGUUCGUGUUCGAAUCCAAAAUGACUACCUCAGUUAAUUAUGGGGCUCCCGAGUAGGGAGC-------------------GAGC--GUCGCGCAUGCGCACAU ----.....((((((.(((((.(((.((((.(((((((.....)))).))).))))((((((.....))))))-------------------....--.))).))))).)))))) ( -34.10, z-score = -2.15, R) >droSim1.chr3R 2692838 90 + 27517382 ----GGGGAGUGUGUUCGUGUUCGAAUCCAAAAUGACUACCUCAGUUAAUUAUGGGGCUCCCGAGUAGGGAGC-------------------GAGC--GUCGCGCAUGCGCACAU ----.....((((((.(((((.(((.((((.(((((((.....)))).))).))))((((((.....))))))-------------------....--.))).))))).)))))) ( -34.10, z-score = -2.15, R) >droSec1.super_6 2749577 90 + 4358794 ----GGGGAGUGUGUUCGUGUUCGAAUCCAAAAUGACUACCUCAGUUAAUUAUGGGGCUCCCGAGUAGGGAGC-------------------GAGC--GUCGCGCAUGCGCACAU ----.....((((((.(((((.(((.((((.(((((((.....)))).))).))))((((((.....))))))-------------------....--.))).))))).)))))) ( -34.10, z-score = -2.15, R) >droYak2.chr3R 21077067 95 - 28832112 ----GGGGAGUGUGUUCGUGUUCGAAUCCAAAAUGACUACCUCAGUUAAUUAUGGGGCUCCCGAGUAGGGAGCCUCCC--------------GAGC--GUCGCGCAUGCGCACAU ----.....((((((.(((((.(((.((.....(((((.....))))).....(((((((((.....)))))))))..--------------))..--.))).))))).)))))) ( -37.80, z-score = -2.54, R) >droEre2.scaffold_4770 2907428 90 + 17746568 ----GGGGAGUGUGUUCGUGUUCGAAUCCAAAAUGACUACCUCAGUUAAUUAUUGGGCUCCCGAGCAGGGAGC-------------------GAAC--GUCGCGCAUGCGCACAU ----.....((((((.(((((.(((..((((..(((((.....)))))....))))((((((.....))))))-------------------....--.))).))))).)))))) ( -33.40, z-score = -2.20, R) >droAna3.scaffold_13340 18302655 90 - 23697760 ----UAGGUGUGUGUUCAUGUUCGAAUCCAAAAUGACUACCUCAGUUAAUUAUGGGGCUCCCGAGUAGGGAGC-------------------GAGC--GUCGCGCAUGCGCACAU ----...((((((((.(((((((...((((.(((((((.....)))).))).))))((((((.....))))))-------------------))))--)).).)))))))).... ( -34.80, z-score = -3.07, R) >dp4.chr2 11082070 98 - 30794189 UGUGUGUAUGCCUGUUCAUGUUCGAAUCCAAAAUGACUACCUCAGUUAAUUAUGGGGCUCCCGAGUAGGGAGCGAGC---------------GAGC--GUCGCGCAUGCGCACAU (((((((((((.((...(((((((..((((.(((((((.....)))).))).))))((((((.....))))))...)---------------))))--)))).))))))))))). ( -42.60, z-score = -4.22, R) >droWil1.scaffold_181089 330970 111 - 12369635 ----UGAGUGCGCGCUCAUGUUCGAAUUCAAAAUGACUACCUCAGUUAAUUAUGGGGCUCCCGAGUAGGGAGCGCACACAUACAAAACAUUCGAGUCAGUCGCGCAUGCGCUUAU ----(((((((((((...(((((((((......(((((.....)))))..((((.(((((((.....)))))).)...))))......))))))).))...))))..))))))). ( -39.20, z-score = -3.20, R) >droGri2.scaffold_14906 2757724 90 - 14172833 ----UGUGUGUGUGUGCAUGUUCGAAUUCAAAAUGACUACCUCAGUUAAUUAUGGGGCUCCCGAGUAGGGAGC-------------------AAGC--GUCGCGCUUGCGCAUAU ----((((((((.(((((((((..................((((........))))((((((.....))))))-------------------.)))--)).)))).)))))))). ( -32.00, z-score = -2.31, R) >consensus ____GGGGAGUGUGUUCGUGUUCGAAUCCAAAAUGACUACCUCAGUUAAUUAUGGGGCUCCCGAGUAGGGAGC___________________GAGC__GUCGCGCAUGCGCACAU .........((((((.(((((.(((.((((.(((((((.....)))).))).))))((((((.....))))))..........................))).))))).)))))) (-29.93 = -29.93 + 0.00)

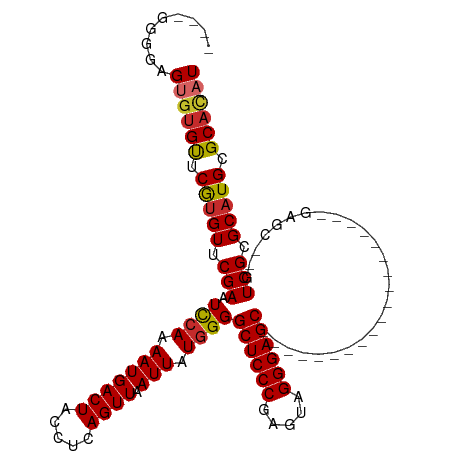
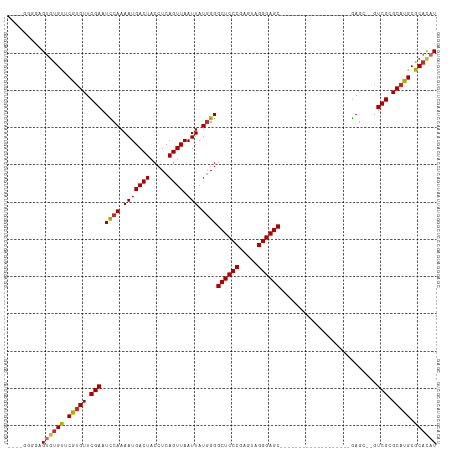
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:54:39 2011