
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 2,489,053 – 2,489,119 |
| Length | 66 |
| Max. P | 0.775516 |

| Location | 2,489,053 – 2,489,119 |
|---|---|
| Length | 66 |
| Sequences | 11 |
| Columns | 77 |
| Reading direction | forward |
| Mean pairwise identity | 72.44 |
| Shannon entropy | 0.52190 |
| G+C content | 0.51584 |
| Mean single sequence MFE | -16.67 |
| Consensus MFE | -8.90 |
| Energy contribution | -8.45 |
| Covariance contribution | -0.45 |
| Combinations/Pair | 1.27 |
| Mean z-score | -1.34 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.65 |
| SVM RNA-class probability | 0.775516 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
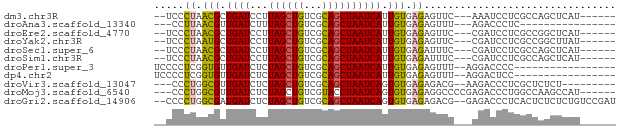
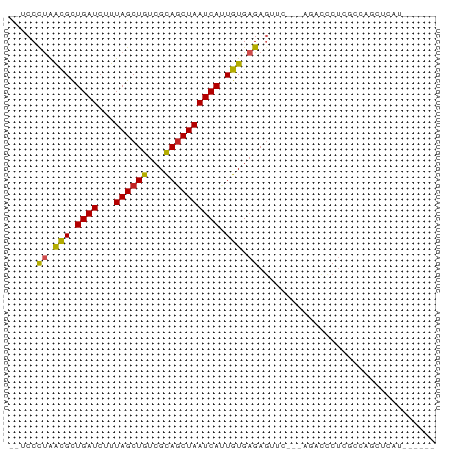
>dm3.chr3R 2489053 66 + 27905053 --UCCCUAACGCUGAUCCUUAGCUGUCGCAGCUAAUCAUUGUGAGAGUUC---AAAUCCUCGCCAGCUCAU------ --........((((....(((((((...))))))).....(((((..(..---..)..)))))))))....------ ( -16.80, z-score = -2.54, R) >droAna3.scaffold_13340 18151911 55 - 23697760 ---CCUUAACGUUGAUCUUUAGCUGUCGCAGCUAAUCAUUGUGAGAGUUU---AGACCCUC---------------- ---............(((..((((.((((((.......)))))).)))).---))).....---------------- ( -10.90, z-score = -1.05, R) >droEre2.scaffold_4770 2732214 66 + 17746568 --UCCCUAACGCUGAUCCUUAGCUGUCGCAGCUAAUCAUUGUGAGAGUUC---CGAUCCUCGCCGGCUCAU------ --........((((....(((((((...))))))).....(((((..(..---..)..)))))))))....------ ( -16.30, z-score = -1.12, R) >droYak2.chr3R 20905119 66 - 28832112 --UCCCUAAUGCUGAUCCUUAGCUGUCGCAGCUAAUCAUUGUGAGAGUUC---CGAUCCUCGCCGGCUUAU------ --........((((....(((((((...))))))).....(((((..(..---..)..)))))))))....------ ( -16.00, z-score = -1.25, R) >droSec1.super_6 2580892 66 + 4358794 --UCCCUAACGCUGAUCCUUAGCUGUCGCAGCUAAUCAUUGUGAGAUUUC---CGAUCCUCGCCAGCUCAU------ --........((((....(((((((...))))))).....(((((.....---.....)))))))))....------ ( -16.50, z-score = -2.22, R) >droSim1.chr3R 2521129 66 + 27517382 --UCCCUAACGCUGAUCCUUAGCUGUCGCAGCUAAUCAUUGUGAGAUUUC---CGAUCCUCGCCAGCUCAU------ --........((((....(((((((...))))))).....(((((.....---.....)))))))))....------ ( -16.50, z-score = -2.22, R) >droPer1.super_3 4741946 58 - 7375914 UCCCCUCGGUGUUGAUCUCUAGCUGUCGCAGCUAAUCAUUGUGAGAGUUU--AGGACCCC----------------- .......(((.((((.(((((((((...)))))).((.....))))).))--)).)))..----------------- ( -15.10, z-score = -1.25, R) >dp4.chr2 10908730 58 - 30794189 UCCCCUCGGUGUUGAUCUCUAGCUGUCGCAGCUAAUCAUUGUGAGAGUUU--AGGACUCC----------------- .......(((.((((.(((((((((...)))))).((.....))))).))--)).)))..----------------- ( -12.70, z-score = -0.05, R) >droVir3.scaffold_13047 12943058 63 - 19223366 ---CCCUGGCGUUGAUCUCUAGCUGUCGCAGCUAAUCAGUGUGAGAGACG--AAGACCCUCGCUCUCU--------- ---.....((((((((...((((((...))))))))))))))(((((.((--(......)))))))).--------- ( -20.20, z-score = -1.29, R) >droMoj3.scaffold_6540 1794622 68 + 34148556 ---CCCUGGCGUUGAUCUCUAGCUGUCGUACCUAAUCAGUGUGAGAGGCCCCGAGACCCUGGCCAAGCCAU------ ---...((((.....((((..((((...........))))..))))((((..(....)..))))..)))).------ ( -19.80, z-score = -1.02, R) >droGri2.scaffold_14906 2570951 73 - 14172833 --CCCCUGGCGAUGAUCUCUAGCUGUCGCAGCUAAUCAGUGUGAGAGACG--GAGACCCUCACUCUCUCUGUCCGAU --...((.(((.((((...((((((...)))))))))).))).)).((((--((((.........)))))))).... ( -22.60, z-score = -0.68, R) >consensus __UCCCUAACGCUGAUCUUUAGCUGUCGCAGCUAAUCAUUGUGAGAGUUC___AGACCCUCGCCAGCUCAU______ .....((.(((.((((...((((((...)))))))))).))).))................................ ( -8.90 = -8.45 + -0.45)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:54:09 2011