
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 2,328,279 – 2,328,376 |
| Length | 97 |
| Max. P | 0.892005 |

| Location | 2,328,279 – 2,328,376 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 116 |
| Reading direction | forward |
| Mean pairwise identity | 86.71 |
| Shannon entropy | 0.22875 |
| G+C content | 0.49202 |
| Mean single sequence MFE | -33.15 |
| Consensus MFE | -25.87 |
| Energy contribution | -26.23 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.05 |
| Mean z-score | -2.08 |
| Structure conservation index | 0.78 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.10 |
| SVM RNA-class probability | 0.892005 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
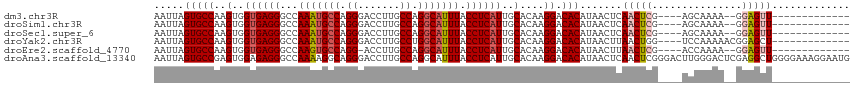
>dm3.chr3R 2328279 97 + 27905053 AAUUAGUGCCAAGUGGUGAGGGCCAAAUGCCAGGGACCUUGCCAGGCAUUUACCUCAUUGCACAAGGACACAUAACUCAACUCG----AGCAAAA--GGAGUU------------- .....(((((..(..((((((...(((((((.((.......)).))))))).))))))..)....)).))).......(((((.----.......--.)))))------------- ( -31.00, z-score = -1.90, R) >droSim1.chr3R 2356869 97 + 27517382 AAUUAGUGCCAAGUGGUGAGGGCCAAAUGCCAGGGACCUUGCCAGGCAUUUACCUCAUUGCACAAGGACACAUAACUCAACUCG----AGCAAAA--GGAGUU------------- .....(((((..(..((((((...(((((((.((.......)).))))))).))))))..)....)).))).......(((((.----.......--.)))))------------- ( -31.00, z-score = -1.90, R) >droSec1.super_6 2414880 97 + 4358794 AAUUAGUGCCAAGUGGUGAGGGCCAAAUGCCAGGGACCUUGCCAGGCAUUUACCUCAUUGCACAAGGACACAUAACUCAACUCG----AGCAAAA--GGAGUU------------- .....(((((..(..((((((...(((((((.((.......)).))))))).))))))..)....)).))).......(((((.----.......--.)))))------------- ( -31.00, z-score = -1.90, R) >droYak2.chr3R 18247892 99 + 28832112 AAUUAGUGCCAAGUGGUGAGGGCCAAAUGCCAGGGACCUUGCCUGGCAUUUACCUCAUUGCACAAGGACACAUAACUUAACUGG----UCCAAAAACGGAGCU------------- ....(((.((..(..((((((...((((((((((.......)))))))))).))))))..)....((((.((.........)))----)))......)).)))------------- ( -38.70, z-score = -3.52, R) >droEre2.scaffold_4770 2574081 96 + 17746568 AAUUAGUGCCAAGUGGUGAGGGCCAAGUGCCAGG-ACCUUGCCAGGCAUUUACCUCAUUGCACAAGGACACAUAACUUAACUCG----ACCAAAA--GGAGUU------------- .....(((((..(..((((((...(((((((.((-......)).))))))).))))))..)....)).))).......(((((.----.......--.)))))------------- ( -31.30, z-score = -2.28, R) >droAna3.scaffold_13340 13555258 116 + 23697760 AAUUAGUGCCGAGUGGAGAGGGCCAAAAGGCAGGGACCUUGCCAGGCAUUUACCUCAUUGCACAAGGACACAUAACUCAACUCGGGACUUGGGACUCGAGGCUGGGGAAAGGAAUG .....(((((..(((.((((((((....((((((...)))))).))).....))))..).)))..)).)))....((((.((((((........))))))..)))).......... ( -35.90, z-score = -0.97, R) >consensus AAUUAGUGCCAAGUGGUGAGGGCCAAAUGCCAGGGACCUUGCCAGGCAUUUACCUCAUUGCACAAGGACACAUAACUCAACUCG____AGCAAAA__GGAGUU_____________ .....(((((..(..((((((...(((((((.((.......)).))))))).))))))..)....)).)))............................................. (-25.87 = -26.23 + 0.36)


| Location | 2,328,279 – 2,328,376 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 86.71 |
| Shannon entropy | 0.22875 |
| G+C content | 0.49202 |
| Mean single sequence MFE | -30.85 |
| Consensus MFE | -25.06 |
| Energy contribution | -25.73 |
| Covariance contribution | 0.67 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.53 |
| Structure conservation index | 0.81 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.654674 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
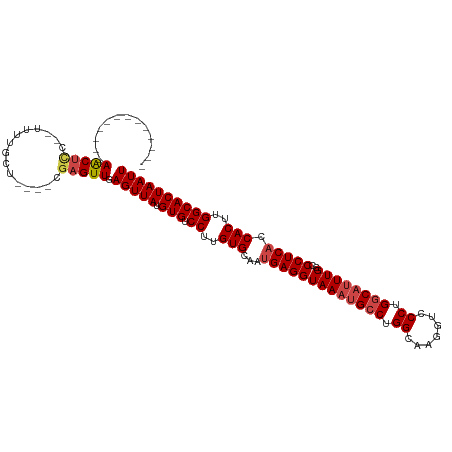
>dm3.chr3R 2328279 97 - 27905053 -------------AACUCC--UUUUGCU----CGAGUUGAGUUAUGUGUCCUUGUGCAAUGAGGUAAAUGCCUGGCAAGGUCCCUGGCAUUUGGCCCUCACCACUUGGCACUAAUU -------------(((((.--.(((...----.)))..)))))..(((.((..(((...(((((((((((((.((.......)).))))))))..))))).)))..)))))..... ( -31.00, z-score = -1.79, R) >droSim1.chr3R 2356869 97 - 27517382 -------------AACUCC--UUUUGCU----CGAGUUGAGUUAUGUGUCCUUGUGCAAUGAGGUAAAUGCCUGGCAAGGUCCCUGGCAUUUGGCCCUCACCACUUGGCACUAAUU -------------(((((.--.(((...----.)))..)))))..(((.((..(((...(((((((((((((.((.......)).))))))))..))))).)))..)))))..... ( -31.00, z-score = -1.79, R) >droSec1.super_6 2414880 97 - 4358794 -------------AACUCC--UUUUGCU----CGAGUUGAGUUAUGUGUCCUUGUGCAAUGAGGUAAAUGCCUGGCAAGGUCCCUGGCAUUUGGCCCUCACCACUUGGCACUAAUU -------------(((((.--.(((...----.)))..)))))..(((.((..(((...(((((((((((((.((.......)).))))))))..))))).)))..)))))..... ( -31.00, z-score = -1.79, R) >droYak2.chr3R 18247892 99 - 28832112 -------------AGCUCCGUUUUUGGA----CCAGUUAAGUUAUGUGUCCUUGUGCAAUGAGGUAAAUGCCAGGCAAGGUCCCUGGCAUUUGGCCCUCACCACUUGGCACUAAUU -------------.(.((((....))))----.).....(((((.(((.((..(((...((((((((((((((((.......)))))))))))..))))).)))..)))))))))) ( -35.80, z-score = -2.51, R) >droEre2.scaffold_4770 2574081 96 - 17746568 -------------AACUCC--UUUUGGU----CGAGUUAAGUUAUGUGUCCUUGUGCAAUGAGGUAAAUGCCUGGCAAGGU-CCUGGCACUUGGCCCUCACCACUUGGCACUAAUU -------------(((((.--.......----.))))).(((((.(((.((..(((...((((((((.((((.((......-)).)))).)))..))))).)))..)))))))))) ( -26.70, z-score = -0.37, R) >droAna3.scaffold_13340 13555258 116 - 23697760 CAUUCCUUUCCCCAGCCUCGAGUCCCAAGUCCCGAGUUGAGUUAUGUGUCCUUGUGCAAUGAGGUAAAUGCCUGGCAAGGUCCCUGCCUUUUGGCCCUCUCCACUCGGCACUAAUU ............((((.(((.(.(....).).)))))))(((((.(((((...(((....((((.....(((.((((.(....)))))....)))))))..)))..)))))))))) ( -29.60, z-score = -0.94, R) >consensus _____________AACUCC__UUUUGCU____CGAGUUGAGUUAUGUGUCCUUGUGCAAUGAGGUAAAUGCCUGGCAAGGUCCCUGGCAUUUGGCCCUCACCACUUGGCACUAAUU .............(((((...............))))).(((((.(((.((..(((...(((((((((((((.((.......)).))))))))..))))).)))..)))))))))) (-25.06 = -25.73 + 0.67)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:53:40 2011