
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 2,121,983 – 2,122,075 |
| Length | 92 |
| Max. P | 0.944983 |

| Location | 2,121,983 – 2,122,075 |
|---|---|
| Length | 92 |
| Sequences | 8 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 81.49 |
| Shannon entropy | 0.34802 |
| G+C content | 0.49580 |
| Mean single sequence MFE | -32.98 |
| Consensus MFE | -18.64 |
| Energy contribution | -18.71 |
| Covariance contribution | 0.07 |
| Combinations/Pair | 1.46 |
| Mean z-score | -2.52 |
| Structure conservation index | 0.57 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.51 |
| SVM RNA-class probability | 0.944983 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 2121983 92 - 27905053 GCUCCGUUUCAUUUGUGCGCUUUUCGG-------UUUGAGAUUUGUUGCUGGUAGUCGCUCAAUUAGCCGGCAACAAGUUGCCAUAA--AUCGAUAGC-CGC ..............(((.(((..((((-------((((.(((((((((((((((((......))).))))))))))))))....)))--))))).)))-))) ( -31.30, z-score = -2.82, R) >droSim1.chr3R 2151463 92 - 27517382 GCUCCGUUUCAUUUGUGCGCUUUUCGG-------UUUGCGAUUUGUUGCUGGCCGUCGCUCAAUUAGCCGGCAACAAGUUGCCAUAA--AUCGAUAGC-CGC ..............(((.(((..((((-------(((((((((((((((((((.............))))))))))))))))...))--))))).)))-))) ( -36.12, z-score = -3.73, R) >droSec1.super_6 2208051 92 - 4358794 GCUCCGUUUCAUUUGUGCGCUUUUCGG-------UUUGCGAUUUGUUGCUGGCCGUCGCUCAAUUAGCUGGCAACAAGUUGCCAUAA--AUCGAUAGC-CGC ..............(((.(((..((((-------(((((((((((((((..((.............))..))))))))))))...))--))))).)))-))) ( -33.52, z-score = -2.92, R) >droEre2.scaffold_4770 2366326 99 - 17746568 GCUCCGUUUUAUUUGUGCGCUUUUCGGUUUGCGAUUUGCGAUUUGCUGCUGGCCAUCGCUCAAUUAGCCGGCAACAAGUUGCCAUAA--AUCGAUAGC-CGC ((.(((.......)).).))....(((((..((((((((((((((.(((((((.............))))))).))))))))...))--))))..)))-)). ( -32.22, z-score = -1.45, R) >droAna3.scaffold_13340 5656901 93 + 23697760 GCUCCAUUUCAUUUGUGCGCUUUUCGA-------UUUGCGAUUUGUUGCUGGGCGUCGCUCAAUUAGCCGGCAACAAGUUGCCAUAA--AUCGAUGGCUCAC ..............(((.(((..((((-------((((((((((((((((((.(............))))))))))))))))...))--))))).))).))) ( -35.10, z-score = -3.65, R) >droYak2.chr3R 18048805 92 - 28832112 GCUCCGUUUUAUUUGUGCGCUUUUCGG-------UUUGUGAUUUGUUGCUGGACGUCGCUCAAUUAGCCGGCAACAAGUUGCCAUAA--AUCGAUAGC-CGC ..............(((.(((..((((-------((((((((((((((((((.(............)))))))))))))...)))))--))))).)))-))) ( -32.00, z-score = -2.92, R) >dp4.chr2 15083881 99 + 30794189 UCUUCGUUUCUUUUUUGCG--ACUCGACUAGCGAUUUGUUGCUGGGUGUCGGGCGUCGCUCAAUUAGCCGGCAACAAGUUGGCCUAAACACCGAUUGU-CGA ...((((.........)))--).(((((...((((((((((((((.((..((((...))))...)).)))))))))))))((........)))...))-))) ( -31.80, z-score = -1.36, R) >droPer1.super_0 4703888 99 - 11822988 UCUUCGUUUCUUUUCUGCG--ACUCGACUAGCGAUUUGUUGCUGGGUGUCGGGCGUCGCUCAAUUAGCCGGCAACAAGUUGGCCUAAACACCGAUUGU-CGA ...((((.........)))--).(((((...((((((((((((((.((..((((...))))...)).)))))))))))))((........)))...))-))) ( -31.80, z-score = -1.28, R) >consensus GCUCCGUUUCAUUUGUGCGCUUUUCGG_______UUUGCGAUUUGUUGCUGGCCGUCGCUCAAUUAGCCGGCAACAAGUUGCCAUAA__AUCGAUAGC_CGC ..............(((.(((..((((..........((((((((.((((((...............)))))).))))))))........)))).))).))) (-18.64 = -18.71 + 0.07)
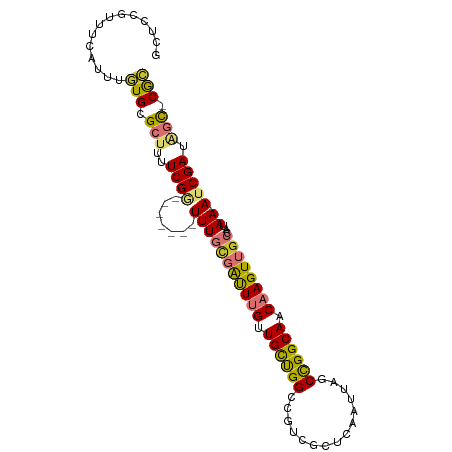
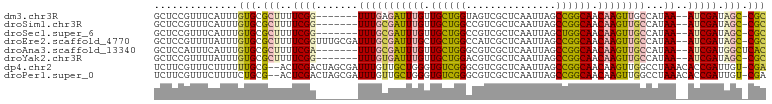

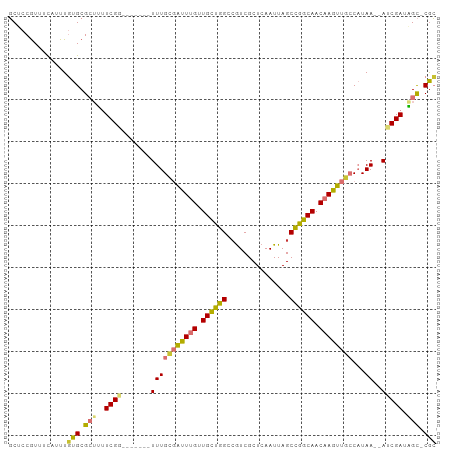
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:53:19 2011