
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 1,829,394 – 1,829,455 |
| Length | 61 |
| Max. P | 0.627926 |

| Location | 1,829,394 – 1,829,455 |
|---|---|
| Length | 61 |
| Sequences | 14 |
| Columns | 61 |
| Reading direction | forward |
| Mean pairwise identity | 80.38 |
| Shannon entropy | 0.46481 |
| G+C content | 0.62061 |
| Mean single sequence MFE | -21.61 |
| Consensus MFE | -12.89 |
| Energy contribution | -11.99 |
| Covariance contribution | -0.90 |
| Combinations/Pair | 2.00 |
| Mean z-score | -1.18 |
| Structure conservation index | 0.60 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.627926 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 1829394 61 + 27905053 CUAGUCCACGGGCCGCCACAUCGGUGGCCACCAGAAUGUUGGACUUGCCCGAGCGGAACUC ....(((.(((((.(((((....)))))..((((....))))....)))))...))).... ( -22.30, z-score = -1.14, R) >apiMel3.GroupUn 251725953 61 - 399230636 CUAAUCCUCUUGCUGCAACAUCUGUAGCUACUAGAAUGGCAGAUCUGCUAUUCCGGAACUC ....(((....((((((.....)))))).....((((((((....)))))))).))).... ( -20.00, z-score = -3.99, R) >anoGam1.chr2R 19467530 61 - 62725911 CCAAACCACGGGCGGCCACAUCUGUUGCUACCAGGAUGCAGGAGCUGGAAUUGCGGAAGCG (((..((...((..((.((....)).))..)).))..((....)))))....((....)). ( -16.40, z-score = 0.46, R) >droGri2.scaffold_14624 1201322 61 - 4233967 CCAAGCCGCGGGCUGCCACAUCGGUGGCAACCAGAAUGUUGGACUUGCCCGAGCGGAACUC .....((((((((((((((....)))))).((((....))))....))))..))))..... ( -25.80, z-score = -2.18, R) >droMoj3.scaffold_6540 14663217 61 - 34148556 CUAUGCCGCGCGCCCACACAUCGGUGGUGAUGAGGACGCGGGACUGGCCGGCACGGAACUC (..((((.(((((((((((.......))).)).)).))))((.....))))))..)..... ( -21.30, z-score = -0.00, R) >droVir3.scaffold_12822 2021450 61 - 4096053 CCAAGCCACGGGCCGCCACAUCGGUGGCCACCAGAAUGUUGGACUUGCCGGAGCGGAAUUC ....((..(((((((((.....))))))).((((....))))......))..))....... ( -22.50, z-score = -1.02, R) >droWil1.scaffold_181130 5197342 61 - 16660200 CCAAACCACGGGCUGCCACAUCGGUGGCUACCAAGAUGUUUGAUUUGCCCGAACGGAAUUC .....((.(((((.(((((....)))))...((((...))))....)))))...))..... ( -18.00, z-score = -1.31, R) >droPer1.super_0 4458009 61 + 11822988 CCAAACCACGGGCCGCCACAUCGGUGGCUACAAGAAUGUUCGAUUUGCCCGAACGGAAUUC .....((...(((((((.....)))))))........(((((.......)))))))..... ( -20.00, z-score = -1.88, R) >dp4.chr2 14840250 61 - 30794189 CCAGACCACGGGCCGCCACAUCGGUGGCUACCAGAAUGUUCGAUUUGCCCGAACGGAAUUC .....((...(((((((.....)))))))........(((((.......)))))))..... ( -20.00, z-score = -1.56, R) >droAna3.scaffold_13340 5413677 61 - 23697760 CCAGGCCUCGGGCUGCCACGUCGGUGGCCACCAGGAUGUUGGACUUGCCGGAACGGAACUC ((.(((....(((..((.....))..))).((((....))))....))))).......... ( -20.20, z-score = 0.71, R) >droEre2.scaffold_4770 2094138 61 + 17746568 CCAGUCCACGGGCCGCCACAUCGGUGGCCACCAGGAUGUUGGACUUUCCCGAGCGGAACUC ((.((((...(((((((.....)))))))....)))).((((......))))..))..... ( -23.50, z-score = -1.20, R) >droYak2.chr3R 17775149 61 + 28832112 CCAGUCCUCGGGCCGCCACAUCGGUGGCCACCAGGAUGUUGGACUUUCCCGAGCGGAACUC ((.(((((.((((((((.....))))))).).))))).((((......))))..))..... ( -25.60, z-score = -1.76, R) >droSec1.super_6 1938503 61 + 4358794 CCAGUCCACGGGCCGCCACAUCGGUGGCCACCAGGAUGUUGGACUUGCCCGAGCGGAACUC ((.((((...(((((((.....)))))))....)))).((((......))))..))..... ( -23.50, z-score = -0.81, R) >droSim1.chr3R_random 218608 61 + 1307089 CCAGUCCACGGGCCGCCACAUCGGUGGCCACCAGGAUGUUGGACUUGCCCGAGCGGAACUC ((.((((...(((((((.....)))))))....)))).((((......))))..))..... ( -23.50, z-score = -0.81, R) >consensus CCAGUCCACGGGCCGCCACAUCGGUGGCCACCAGGAUGUUGGACUUGCCCGAGCGGAACUC .....((...(((((((.....)))))))........(((...........)))))..... (-12.89 = -11.99 + -0.90)


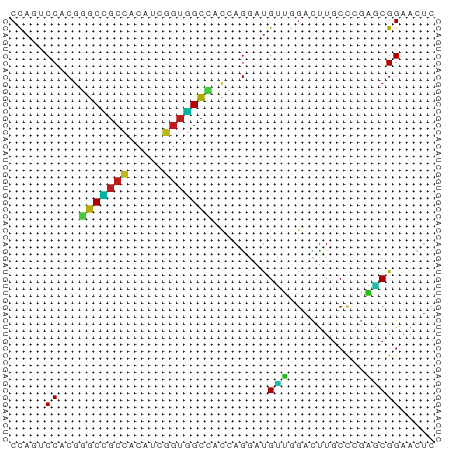
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:52:42 2011