
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 1,751,353 – 1,751,451 |
| Length | 98 |
| Max. P | 0.832012 |

| Location | 1,751,353 – 1,751,451 |
|---|---|
| Length | 98 |
| Sequences | 9 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 82.64 |
| Shannon entropy | 0.35824 |
| G+C content | 0.34977 |
| Mean single sequence MFE | -22.13 |
| Consensus MFE | -11.58 |
| Energy contribution | -12.22 |
| Covariance contribution | 0.65 |
| Combinations/Pair | 1.26 |
| Mean z-score | -2.31 |
| Structure conservation index | 0.52 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.84 |
| SVM RNA-class probability | 0.832012 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
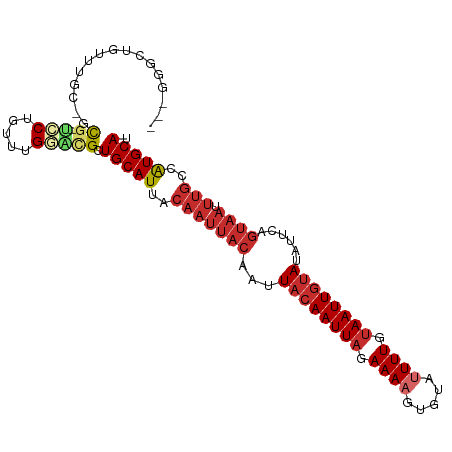
>dm3.chr3R 1751353 98 + 27905053 AUGCAUGGCAAAUUACUGAAUAUACAAUUACAAAAUACACUUUUCUAAUUGUAAUUGUAAUUGUAAUGCAGCGUCCAAACAGGA-CGG-GCAAACAACCC--- .(((...........(((....(((((((((((..((((.((....)).)))).)))))))))))...)))(((((.....)))-)).-)))........--- ( -24.80, z-score = -3.23, R) >droEre2.scaffold_4770 2018157 98 + 17746568 AUGCAUGGCAAAUUACUGAAUAUACAAUUACAAAAUACACUUUUCUAAUUGUAAUUGUAAUUGUAAUGCAGCGCCCAAACAGGA-CGG-GCAAACAGCCC--- ......(((......(((....(((((((((((..((((.((....)).)))).)))))))))))...))).((((........-.))-)).....))).--- ( -25.80, z-score = -2.94, R) >droSec1.super_6 1859846 98 + 4358794 AUGCAUGGCAAAUUACUGAAUAUACAAUUACAAAAUACACUUUUCUAAUUGUAAUUGUAAUUGUAAUGCAGCGUACAAACAGGA-CGG-GCAAACAACCC--- .(((((.((((.((((......((((((((.((((.....)))).))))))))...)))))))).)))))..............-.((-(.......)))--- ( -18.50, z-score = -1.11, R) >droSim1.chr3R 1786398 98 + 27517382 AUGCAUGGCAAAUUACUGAAUAUACAAUUACAAAACACACUUUUCUAAUUGUAAUUGUAAUUGUAAUGCAGCGUCCAAACAGGA-CGG-GCAAACAACCC--- .(((((.((((.((((......((((((((.((((.....)))).))))))))...)))))))).)))))..((((.....)))-)((-(.......)))--- ( -24.20, z-score = -2.97, R) >droAna3.scaffold_13340 5342088 98 - 23697760 AUGCAUGCCAAAUUACUGAAUAUACAAUUACAAAAUACACUUUUCUAAUUGUAAUUGUAAUUGUAAUGCAGCGUCCAAACAGGA-GGC-ACAACAAGCAA--- .(((.(((((......))....(((((((((((..((((.((....)).)))).)))))))))))..)))((.(((.....)))-.))-.......))).--- ( -23.90, z-score = -2.98, R) >dp4.chr2 14762301 96 - 30794189 AUGCACAGCAAAUUACGGAAUAUACAAUUACAAAGUACACUUUUCUAAUUGUAAUUGUAAUUGUAAUGCAGCGGGCAAGCAGCC-CACAGCAAACAG------ .(((...((..(((((((....(((((((((((((.........))..))))))))))).)))))))...))((((.....)))-)...))).....------ ( -25.80, z-score = -3.41, R) >droPer1.super_0 4384412 96 + 11822988 AUGCACAGCAAAUUACGGAAUAUACAAUUACAAAGUACACUUUUCUAAUUGUAAUUGUAAUUGUAAUGCAGCGGCCAAGCAGCC-CACAGCAAACAG------ .(((...((..(((((((....(((((((((((((.........))..))))))))))).)))))))...))(((......)))-....))).....------ ( -21.30, z-score = -1.93, R) >droWil1.scaffold_181130 5111291 97 - 16660200 AUGCACAGCAAAUUACUGAAUCUACAAUUACAAAAUACACUUUUCUAAUUGUAAUUGUAAUUGUAAUGCAACAGCCAAACAGGAGCACAGCAAACAG------ .(((.(((.......)))....(((((((((((..((((.((....)).)))).))))))))))).(((.....((.....)).)))..))).....------ ( -17.60, z-score = -1.62, R) >droGri2.scaffold_14624 1119383 90 - 4233967 AUGCAUG------CAACGUGUAUGUAAUUACAAAAUACACUUUUCAAAUUGUAAUU------GCAAUGCAACUGUCAAAAUGGG-GACUGUAAACAAACCGAA .((((((------(.....))))))).........((((.((..((..(((((.((------((...)))).)).)))..))..-)).))))........... ( -17.30, z-score = -0.56, R) >consensus AUGCAUGGCAAAUUACUGAAUAUACAAUUACAAAAUACACUUUUCUAAUUGUAAUUGUAAUUGUAAUGCAGCGGCCAAACAGGA_CGC_GCAAACAGCCC___ ......................(((((((((((..((((.((....)).)))).))))))))))).(((....(((.....))).....)))........... (-11.58 = -12.22 + 0.65)



| Location | 1,751,353 – 1,751,451 |
|---|---|
| Length | 98 |
| Sequences | 9 |
| Columns | 103 |
| Reading direction | reverse |
| Mean pairwise identity | 82.64 |
| Shannon entropy | 0.35824 |
| G+C content | 0.34977 |
| Mean single sequence MFE | -25.30 |
| Consensus MFE | -13.11 |
| Energy contribution | -13.92 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.30 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.52 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.17 |
| SVM RNA-class probability | 0.577397 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 1751353 98 - 27905053 ---GGGUUGUUUGC-CCG-UCCUGUUUGGACGCUGCAUUACAAUUACAAUUACAAUUAGAAAAGUGUAUUUUGUAAUUGUAUAUUCAGUAAUUUGCCAUGCAU ---((((.....))-))(-(((.....))))((((...(((((((((((.((((.((....)).))))..)))))))))))....))))....(((...))). ( -28.90, z-score = -3.26, R) >droEre2.scaffold_4770 2018157 98 - 17746568 ---GGGCUGUUUGC-CCG-UCCUGUUUGGGCGCUGCAUUACAAUUACAAUUACAAUUAGAAAAGUGUAUUUUGUAAUUGUAUAUUCAGUAAUUUGCCAUGCAU ---((((.....))-))(-(((.....))))((((...(((((((((((.((((.((....)).))))..)))))))))))....))))....(((...))). ( -30.40, z-score = -3.03, R) >droSec1.super_6 1859846 98 - 4358794 ---GGGUUGUUUGC-CCG-UCCUGUUUGUACGCUGCAUUACAAUUACAAUUACAAUUAGAAAAGUGUAUUUUGUAAUUGUAUAUUCAGUAAUUUGCCAUGCAU ---((((.....))-)).-...(((..(((.((((...(((((((((((.((((.((....)).))))..)))))))))))....))))....)))...))). ( -23.30, z-score = -1.66, R) >droSim1.chr3R 1786398 98 - 27517382 ---GGGUUGUUUGC-CCG-UCCUGUUUGGACGCUGCAUUACAAUUACAAUUACAAUUAGAAAAGUGUGUUUUGUAAUUGUAUAUUCAGUAAUUUGCCAUGCAU ---((((.....))-))(-(((.....))))((((...(((((((((((.((((.((....)).))))..)))))))))))....))))....(((...))). ( -28.60, z-score = -2.90, R) >droAna3.scaffold_13340 5342088 98 + 23697760 ---UUGCUUGUUGU-GCC-UCCUGUUUGGACGCUGCAUUACAAUUACAAUUACAAUUAGAAAAGUGUAUUUUGUAAUUGUAUAUUCAGUAAUUUGGCAUGCAU ---.(((.....((-(((-(((.....))).((((...(((((((((((.((((.((....)).))))..)))))))))))....)))).....)))))))). ( -26.80, z-score = -2.50, R) >dp4.chr2 14762301 96 + 30794189 ------CUGUUUGCUGUG-GGCUGCUUGCCCGCUGCAUUACAAUUACAAUUACAAUUAGAAAAGUGUACUUUGUAAUUGUAUAUUCCGUAAUUUGCUGUGCAU ------.........(((-(((.....))))))(((((..(((((((...((((((((..((((....)))).))))))))......)))).)))..))))). ( -28.20, z-score = -2.84, R) >droPer1.super_0 4384412 96 - 11822988 ------CUGUUUGCUGUG-GGCUGCUUGGCCGCUGCAUUACAAUUACAAUUACAAUUAGAAAAGUGUACUUUGUAAUUGUAUAUUCCGUAAUUUGCUGUGCAU ------.........(((-(.(.....).))))(((((..(((((((...((((((((..((((....)))).))))))))......)))).)))..))))). ( -23.60, z-score = -0.78, R) >droWil1.scaffold_181130 5111291 97 + 16660200 ------CUGUUUGCUGUGCUCCUGUUUGGCUGUUGCAUUACAAUUACAAUUACAAUUAGAAAAGUGUAUUUUGUAAUUGUAGAUUCAGUAAUUUGCUGUGCAU ------......((((.((....)).))))...((((((((((((((((.((((.((....)).))))..))))))))))))...((((.....)))))))). ( -22.70, z-score = -1.10, R) >droGri2.scaffold_14624 1119383 90 + 4233967 UUCGGUUUGUUUACAGUC-CCCAUUUUGACAGUUGCAUUGC------AAUUACAAUUUGAAAAGUGUAUUUUGUAAUUACAUACACGUUG------CAUGCAU ...((.(((....))).)-).............(((((.((------(((..(.....)....(((((...(((....))))))))))))------)))))). ( -15.20, z-score = 0.26, R) >consensus ___GGGCUGUUUGC_GCG_UCCUGUUUGGACGCUGCAUUACAAUUACAAUUACAAUUAGAAAAGUGUAUUUUGUAAUUGUAUAUUCAGUAAUUUGCCAUGCAU ...........(((.....(((.....))).((((...(((((((((((.((((.((....)).))))..)))))))))))....))))..........))). (-13.11 = -13.92 + 0.81)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:52:30 2011