
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 1,749,300 – 1,749,393 |
| Length | 93 |
| Max. P | 0.815580 |

| Location | 1,749,300 – 1,749,393 |
|---|---|
| Length | 93 |
| Sequences | 10 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 77.96 |
| Shannon entropy | 0.41759 |
| G+C content | 0.47460 |
| Mean single sequence MFE | -20.18 |
| Consensus MFE | -14.15 |
| Energy contribution | -14.56 |
| Covariance contribution | 0.41 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.34 |
| Structure conservation index | 0.70 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.78 |
| SVM RNA-class probability | 0.815580 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 1749300 93 + 27905053 ------ACACGCGC-CUGAGAUUACGCUUUAUGAUACGUGAUAACGCAU--UGCGUAACCUUUU-CGCAGAUAAUCCUGUUAGACUGC-CUACGCACUUAUCCC-------------- ------....(((.-..(((.((((((...(((...(((....))))))--.)))))).)))..-))).(((((...((((((.....-))).))).)))))..-------------- ( -19.70, z-score = -1.76, R) >droSec1.super_6 1857341 93 + 4358794 ------CCACGCGC-CUGAGAUUACGCUUUAUGAUACGUGAUAACGCAU--UGCGUAACCUUUU-CGCAGAUAAUCCUGUUAGACUGC-CUACGCACUUAUCCC-------------- ------....(((.-..(((.((((((...(((...(((....))))))--.)))))).)))..-))).(((((...((((((.....-))).))).)))))..-------------- ( -19.70, z-score = -1.86, R) >droYak2.chr3R 17694281 94 + 28832112 ------CCACGCGC-CUGAGAUUACGCUUUAUGAUACGUGAUAACGCAU--UUCGUAGCCUUUUUCGCAGAUAAUCCUGUUAGACUGC-CUACGCACUUAUCCC-------------- ------....((((-.....((((((..........))))))...))..--...((((.(..(((.((((......)))).)))..).-)))))).........-------------- ( -17.20, z-score = -0.76, R) >droEre2.scaffold_4770 2015821 77 + 17746568 ------CCACGCGC-CUGAGAUUACGCUUUAUGAUACGUGAUAACGCAU--UGCGUAACCUUUU-CGCAGAUAAUCCUGUUAGACUC------------------------------- ------....(((.-..(((.((((((...(((...(((....))))))--.)))))).)))..-)))...................------------------------------- ( -16.40, z-score = -1.38, R) >droAna3.scaffold_13340 5339790 93 - 23697760 ------CCACGCGC-UUGAGAUUACGCUUUAUGAUACGUGAUAACGCAU--UGCGUAACCUUUU-CGCAGAUAAUCCUGUUAGACGGC-CUACGCACUUAUCCU-------------- ------....(((.-.....((((((..........))))))..)))..--((((((.((.(((-.((((......)))).))).)).-.))))))........-------------- ( -20.80, z-score = -1.28, R) >dp4.chr2 14759809 107 - 30794189 ------CCACGCGC-UUGAGAUUACGCUUUAUGAUACGUGAUAACGCAU--UGCGUAACCUUUU-CGCAGAUAAUCUUGUUAGGCGGU-GUACGCCCAUAGAUGCCAUCUCAUCCCCC ------....(((.-..(((.((((((...(((...(((....))))))--.)))))).)))..-)))((((.((((.....((((..-...))))...))))...))))........ ( -27.10, z-score = -1.44, R) >droPer1.super_0 4381966 93 + 11822988 ------CCACGCGC-UUGAGAUUACGCUUUAUGAUACGUGAUAACGCAU--UGCGUAACCUUUU-CGCAGAUAAUCCUGUUAGGCGGU-GUACGCCCAUCCCCC-------------- ------....(((.-..(((.((((((...(((...(((....))))))--.)))))).)))..-)))..............((((..-...))))........-------------- ( -23.30, z-score = -1.23, R) >droWil1.scaffold_181130 5108077 96 - 16660200 ------CCACGCGC-UUGAGAUUACGCUUUAUGAUACGUGAUAACGCAU--UGCGUAACCUUUU-CGCAGAUAAUCCUGUUAGGCGGCACCAAACACCGAACACCG------------ ------....(((.-..(((.((((((...(((...(((....))))))--.)))))).)))..-))).........((((.((.(........).)).))))...------------ ( -21.10, z-score = -0.89, R) >droGri2.scaffold_14624 1116537 99 - 4233967 --CGCACCACGCAC-UUGAGAUUACGCUUUAUGUUACGUGAUAACGCAUAACGCGUAACCUUUU-CACAGAUAAUCUCCUUAAGCAGCGACAGCGCCUACGCC--------------- --.......(((.(-(((((.((((((.(((((...(((....)))))))).))))))......-...((.....)).))))))..)))...(((....))).--------------- ( -22.30, z-score = -1.49, R) >droVir3.scaffold_12822 1925532 87 - 4096053 CCCAUCCCACGCGCCUUCAGAUUACGCUUUAUGUUACGUGAUAACGCAUAACGCGUAACCUUUU-CACAGAUAAUCUCCUUAAGCAGC------------------------------ ..........((......(((((((.......((((((((.((.....)).)))))))).....-....).))))))......))...------------------------------ ( -14.19, z-score = -1.30, R) >consensus ______CCACGCGC_UUGAGAUUACGCUUUAUGAUACGUGAUAACGCAU__UGCGUAACCUUUU_CGCAGAUAAUCCUGUUAGACGGC_CUACGCACUUAUCCC______________ ..........(((....(((.((((((...(((...(((....))))))...)))))).)))...))).................................................. (-14.15 = -14.56 + 0.41)


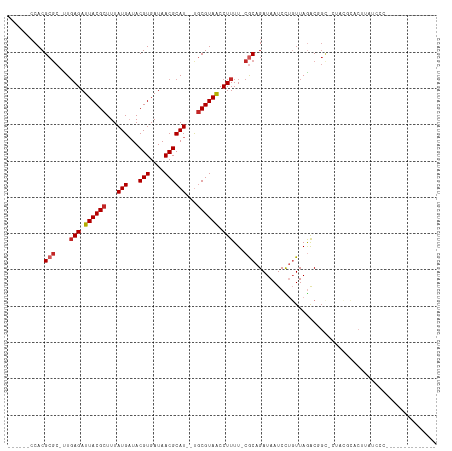
| Location | 1,749,300 – 1,749,393 |
|---|---|
| Length | 93 |
| Sequences | 10 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 77.96 |
| Shannon entropy | 0.41759 |
| G+C content | 0.47460 |
| Mean single sequence MFE | -26.75 |
| Consensus MFE | -17.58 |
| Energy contribution | -17.73 |
| Covariance contribution | 0.15 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.36 |
| Structure conservation index | 0.66 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.58 |
| SVM RNA-class probability | 0.748785 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 1749300 93 - 27905053 --------------GGGAUAAGUGCGUAG-GCAGUCUAACAGGAUUAUCUGCG-AAAAGGUUACGCA--AUGCGUUAUCACGUAUCAUAAAGCGUAAUCUCAG-GCGCGUGU------ --------------.......((((((((-..(((((....)))))..)))).-...(((((((((.--((((((....))))))......)))))))))...-))))....------ ( -29.30, z-score = -2.67, R) >droSec1.super_6 1857341 93 - 4358794 --------------GGGAUAAGUGCGUAG-GCAGUCUAACAGGAUUAUCUGCG-AAAAGGUUACGCA--AUGCGUUAUCACGUAUCAUAAAGCGUAAUCUCAG-GCGCGUGG------ --------------.......((((((((-..(((((....)))))..)))).-...(((((((((.--((((((....))))))......)))))))))...-))))....------ ( -29.30, z-score = -2.79, R) >droYak2.chr3R 17694281 94 - 28832112 --------------GGGAUAAGUGCGUAG-GCAGUCUAACAGGAUUAUCUGCGAAAAAGGCUACGAA--AUGCGUUAUCACGUAUCAUAAAGCGUAAUCUCAG-GCGCGUGG------ --------------..((((.((((((((-..(((((....)))))..)))))...((.((......--..)).))..))).)))).....((((........-))))....------ ( -24.30, z-score = -0.96, R) >droEre2.scaffold_4770 2015821 77 - 17746568 -------------------------------GAGUCUAACAGGAUUAUCUGCG-AAAAGGUUACGCA--AUGCGUUAUCACGUAUCAUAAAGCGUAAUCUCAG-GCGCGUGG------ -------------------------------.(((((....)))))....(((-...(((((((((.--((((((....))))))......)))))))))...-.)))....------ ( -20.70, z-score = -1.75, R) >droAna3.scaffold_13340 5339790 93 + 23697760 --------------AGGAUAAGUGCGUAG-GCCGUCUAACAGGAUUAUCUGCG-AAAAGGUUACGCA--AUGCGUUAUCACGUAUCAUAAAGCGUAAUCUCAA-GCGCGUGG------ --------------.......((((((((-(..((((....))))..))))).-...(((((((((.--((((((....))))))......)))))))))...-))))....------ ( -27.30, z-score = -1.79, R) >dp4.chr2 14759809 107 + 30794189 GGGGGAUGAGAUGGCAUCUAUGGGCGUAC-ACCGCCUAACAAGAUUAUCUGCG-AAAAGGUUACGCA--AUGCGUUAUCACGUAUCAUAAAGCGUAAUCUCAA-GCGCGUGG------ ........(((((..((((.((((((...-..))))))...)))))))))(((-...(((((((((.--((((((....))))))......)))))))))...-.)))....------ ( -31.20, z-score = -1.17, R) >droPer1.super_0 4381966 93 - 11822988 --------------GGGGGAUGGGCGUAC-ACCGCCUAACAGGAUUAUCUGCG-AAAAGGUUACGCA--AUGCGUUAUCACGUAUCAUAAAGCGUAAUCUCAA-GCGCGUGG------ --------------......((((((...-..))))))............(((-...(((((((((.--((((((....))))))......)))))))))...-.)))....------ ( -26.90, z-score = -1.02, R) >droWil1.scaffold_181130 5108077 96 + 16660200 ------------CGGUGUUCGGUGUUUGGUGCCGCCUAACAGGAUUAUCUGCG-AAAAGGUUACGCA--AUGCGUUAUCACGUAUCAUAAAGCGUAAUCUCAA-GCGCGUGG------ ------------...((((.((((........)))).)))).........(((-...(((((((((.--((((((....))))))......)))))))))...-.)))....------ ( -25.90, z-score = -0.14, R) >droGri2.scaffold_14624 1116537 99 + 4233967 ---------------GGCGUAGGCGCUGUCGCUGCUUAAGGAGAUUAUCUGUG-AAAAGGUUACGCGUUAUGCGUUAUCACGUAACAUAAAGCGUAAUCUCAA-GUGCGUGGUGCG-- ---------------.......((((((.(((.((((..(((.....)))...-...(((((((((.((((((((....)))...))))).))))))))).))-))))))))))).-- ( -30.10, z-score = -0.55, R) >droVir3.scaffold_12822 1925532 87 + 4096053 ------------------------------GCUGCUUAAGGAGAUUAUCUGUG-AAAAGGUUACGCGUUAUGCGUUAUCACGUAACAUAAAGCGUAAUCUGAAGGCGCGUGGGAUGGG ------------------------------.(..(....(((.....)))(((-...(((((((((.((((((((....)))...))))).))))))))).....))))..)...... ( -22.50, z-score = -0.77, R) >consensus ______________GGGAUAAGUGCGUAG_GCAGCCUAACAGGAUUAUCUGCG_AAAAGGUUACGCA__AUGCGUUAUCACGUAUCAUAAAGCGUAAUCUCAA_GCGCGUGG______ ..................................................(((....(((((((((...((((((....))))))......))))))))).....))).......... (-17.58 = -17.73 + 0.15)



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:52:28 2011