
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 1,545,298 – 1,545,362 |
| Length | 64 |
| Max. P | 0.654233 |

| Location | 1,545,298 – 1,545,362 |
|---|---|
| Length | 64 |
| Sequences | 12 |
| Columns | 70 |
| Reading direction | reverse |
| Mean pairwise identity | 72.33 |
| Shannon entropy | 0.57462 |
| G+C content | 0.53856 |
| Mean single sequence MFE | -22.06 |
| Consensus MFE | -7.38 |
| Energy contribution | -7.77 |
| Covariance contribution | 0.38 |
| Combinations/Pair | 1.43 |
| Mean z-score | -1.95 |
| Structure conservation index | 0.33 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.654233 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 1545298 64 - 27905053 CUUGACCAGCUCUAAACGGCUGAGAUUACUCGGUCGCGUGUCCAU------AGUGGGGUUGGUCAGUCCG .(((((((((((((..((((((((....)))))))).........------..))))))))))))).... ( -27.22, z-score = -3.42, R) >droSim1.chr3R 1582164 64 - 27517382 CUUGACCAGCUCUAAACGGCUGAGAUUACUCGGUCGCGUGUCCAU------CGUGGGGUUGGUCAGUCCG .(((((((((((((..((((((((....))))))))((.......------))))))))))))))).... ( -28.20, z-score = -3.63, R) >droSec1.super_6 1660916 64 - 4358794 CUUGACCAGCUCUAAACGGCUGAGAUUACUCGGUCGCGUGUCCAU------CGUGGGGUUGGUCAGUCCG .(((((((((((((..((((((((....))))))))((.......------))))))))))))))).... ( -28.20, z-score = -3.63, R) >droYak2.chr3R 17509001 63 - 28832112 CUUGACCAGUUCC-AGCGGCUGAGAUUACUCGGACGCGUGUCCAU------CGUGGGGUUGGUCAGUUCG .((((((((..((-(.(((..(((....)))((((....)))).)------)))))..)))))))).... ( -27.10, z-score = -2.93, R) >droEre2.scaffold_4770 1819609 63 - 17746568 CUUGACCAGUUCU-AACUGCUGGGAUUACUCGGACGCGUGUCCAU------CGUGGGGUUGGUCAGUCCG .((((((((..((-(......(((....)))((((....))))..------..)))..)))))))).... ( -21.80, z-score = -1.30, R) >droAna3.scaffold_13340 5149068 64 + 23697760 CUUGACCAGCUCUGGCUGAAGGAGGUUGCUAGGACACGCGUCCAU------CGCGGGGUUGGUCAGUCGA .(((((((((((((.(.((.(((.((((......)).)).))).)------))))))))))))))).... ( -26.40, z-score = -1.69, R) >dp4.chr2 17384057 63 - 30794189 CUUGACCAGCGAUAAUCGAUCAAAG-CACUCGGACGCGUGUCAUU------CGCGGGGUUGGUCAGUCGA .(((((((((.....((((......-...)))).(.((((.....------)))).)))))))))).... ( -19.60, z-score = -0.71, R) >droPer1.super_0 591570 63 + 11822988 CUUGACCAGCGCUAAUCGAUCAAAG-CACUCGGACGCGUGUCAUU------CGCGGGGUUGGUCAGUCGA .(((((((((.(...((((......-...)))).((((.......------))))).))))))))).... ( -20.70, z-score = -0.94, R) >droWil1.scaffold_181130 8881979 69 - 16660200 CUUGACCGACACAAAAAUAUUUA-AUGACUUAAGCGCGUGUCCAAGCUCAAUGUCCAACUGGUCAGUCAA .(((((((((((...........-.............)))))...((.....))......)))))).... ( -10.76, z-score = -0.11, R) >droVir3.scaffold_13047 12650745 63 + 19223366 AUUGACCGGCAUUAAAUUAUUUA-AUGACUCGGACGCGUGUCCUU------UGCCGGGUUGGUCAGUCAG ((((((((.((((((.....)))-)))((((((.((.........------))))))))))))))))... ( -17.50, z-score = -1.32, R) >droMoj3.scaffold_6540 1509971 63 - 34148556 CUUAACCAGCAUUAAAUGAUUUA-AUGGCUACGACGCGUGUCCAG------UGUCGUAGUGGUCACUCAA ....(((..((((((.....)))-)))((((((((((.......)------))))))))))))....... ( -20.90, z-score = -3.38, R) >droGri2.scaffold_14906 11863562 63 - 14172833 CUUGACCAGCGUUAAAUGAUUUA-AUGACUCAGACGCGUGUCCUU------AGCGGGGUUGGUCAGUGCG .(((((((((((((.........-.)))).....(((........------.)))..))))))))).... ( -16.30, z-score = -0.40, R) >consensus CUUGACCAGCUCUAAACGAUUGAGAUUACUCGGACGCGUGUCCAU______CGUGGGGUUGGUCAGUCCG .(((((((((.((..............((.(......).))..............))))))))))).... ( -7.38 = -7.77 + 0.38)

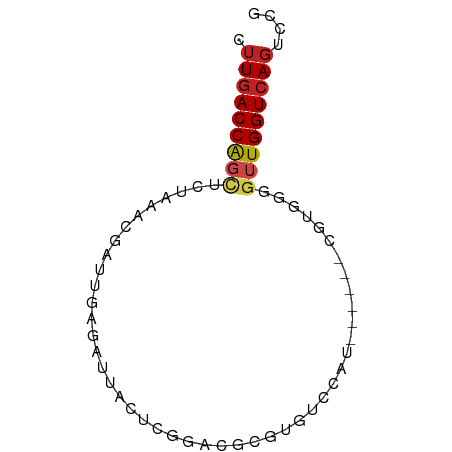

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:52:01 2011