
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 1,438,158 – 1,438,249 |
| Length | 91 |
| Max. P | 0.868857 |

| Location | 1,438,158 – 1,438,249 |
|---|---|
| Length | 91 |
| Sequences | 9 |
| Columns | 103 |
| Reading direction | forward |
| Mean pairwise identity | 74.38 |
| Shannon entropy | 0.48646 |
| G+C content | 0.44411 |
| Mean single sequence MFE | -24.99 |
| Consensus MFE | -13.75 |
| Energy contribution | -13.23 |
| Covariance contribution | -0.52 |
| Combinations/Pair | 1.56 |
| Mean z-score | -1.61 |
| Structure conservation index | 0.55 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.99 |
| SVM RNA-class probability | 0.868857 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 1438158 91 + 27905053 UUUCCGCGCCGAGU-CGUCCCUAAAUAAAUGGCAUAUAUACCGCUGUGCGUGUGUGC--AUAUGUAUAUGUGC-CUCGUCUGUAUGCAGCGUGGU-------- ...((((((.....-...............(((((((((((...((((((....)))--))).))))))))))-)..((......)).)))))).-------- ( -29.30, z-score = -1.54, R) >droSim1.chr3R 1473525 91 + 27517382 UUUCCGCGCCGAGU-CGUCCCUAAAUAAAUGGCAUAUAUACCGCUGUGCGUGUGUAC--AUAUGUAUAUGUGC-CUCGUCUGUAUGCAGCGUGGU-------- ...((((((.....-...............(((((((((((...(((((....))))--)...))))))))))-)..((......)).)))))).-------- ( -29.20, z-score = -1.79, R) >droSec1.super_6 1553398 91 + 4358794 UUUCCGCGCCGAGU-CGUCCCUAAAUAAAUGGCAUAUAUACCGCUGUGCGUUUGUAC--AUAUGUAUAUGUGC-CUCGUCUGUAUGCAGCGUGGU-------- ...((((((.....-...............(((((((((((...(((((....))))--)...))))))))))-)..((......)).)))))).-------- ( -29.20, z-score = -2.27, R) >droYak2.chr3R 17403580 93 + 28832112 UUUCCGCGCCGAGU-CGUACCUAAAUAAAUGGCAUAUAUACCGCUGUGCGUGUAUAUAUGUAUGUAUAUGUGC-CUCGUCUGUAUGCAGCGUGGU-------- ...((((((((((.-.((((.((.(((....(((((((((((((...))).)))))))))).))).)).))))-))))..((....)))))))).-------- ( -30.80, z-score = -1.85, R) >droEre2.scaffold_4770 1709244 91 + 17746568 UUUCCGCGCCGAGU-UGUCCCUAAAUAAAUGGCAUAUAUACCGCUGUAGGUGUGUAC--AUAUGUAUAUGUGC-CUCGUCUGUAUGCAGCGUGGU-------- ...(((((((((.(-(((......))))..(((((((((((...((((......)))--)...))))))))))-))))..((....)))))))).-------- ( -26.40, z-score = -0.98, R) >droAna3.scaffold_13340 5072007 75 - 23697760 UUUCCGCGCCGUAU-CGUUCCUAAAUAAAUGAUAUAGGUAU------------------AUAUAAGUAUAUGC-CUCGUCUGUAUGCAGCGUGCU-------- .....((((.((((-((((........))))))))((((((------------------(((....)))))))-)).(.(((....)))))))).-------- ( -20.80, z-score = -2.77, R) >dp4.chr2 17303856 75 + 30794189 UUUCCGCGCCGUGU-CGUCCCUAAAUAUAUUGCAUAUAUAU------------------GUAUGUGUAUAUGC-UUCGUCUGUAUGCAGCGUGGU-------- ...((((((.((((-((..(...........((((((((((------------------....))))))))))-...)..)).)))).)))))).-------- ( -20.34, z-score = -1.74, R) >droPer1.super_0 11149039 75 + 11822988 UUUCCGCGCCGUGU-CGUCCCUAAAUAUAUUGCAUAUAUAU------------------GUAUGUGUAUAUGC-UUCGUCUGUAUGCAGCGUGGU-------- ...((((((.((((-((..(...........((((((((((------------------....))))))))))-...)..)).)))).)))))).-------- ( -20.34, z-score = -1.74, R) >droGri2.scaffold_15074 4882080 100 + 7742996 UUUCCGCGCCGUCUUCGUCCCUAAAUAUAUUGCGUAUAUAUAUACA---AUAUACAUAUAUAUGUAUGUUCGCACACACACACGCGCGCUGUAUUUGUGUAUU ....(((((((....)).............(((((((....))))(---((((((((....))))))))).))).........)))))............... ( -18.50, z-score = 0.20, R) >consensus UUUCCGCGCCGAGU_CGUCCCUAAAUAAAUGGCAUAUAUACCGCUGU__GUGUGUAC__AUAUGUAUAUGUGC_CUCGUCUGUAUGCAGCGUGGU________ ...((((((......................((((((((((......................)))))))))).......((....))))))))......... (-13.75 = -13.23 + -0.52)


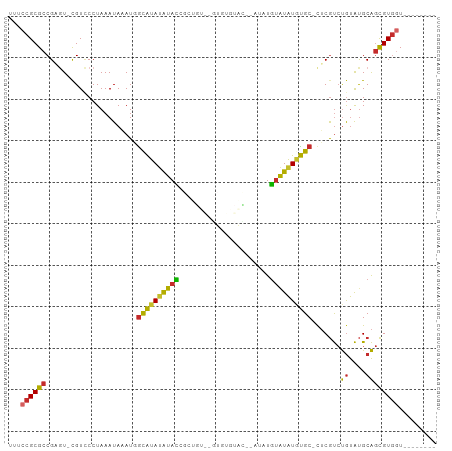
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:51:47 2011