
| Sequence ID | dm3.chr3R |
|---|---|
| Location | 1,297,230 – 1,297,402 |
| Length | 172 |
| Max. P | 0.791341 |

| Location | 1,297,230 – 1,297,330 |
|---|---|
| Length | 100 |
| Sequences | 3 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 89.24 |
| Shannon entropy | 0.14455 |
| G+C content | 0.30210 |
| Mean single sequence MFE | -19.47 |
| Consensus MFE | -15.94 |
| Energy contribution | -16.50 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.82 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.70 |
| SVM RNA-class probability | 0.791341 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
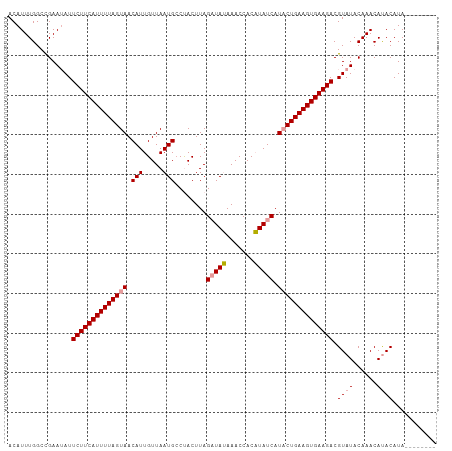
>dm3.chr3R 1297230 100 + 27905053 ACAUUUGGCUGAAGAUUCUUCAUUUUAGUAACAUUGUUAAUGACCACUUAGUUAAAAACCACUUAUCAUUCUGAAGUGAAGACGUAUACAAACACACAUA-------- .....((..((...((((((((((((((((((...)))).((((......))))................)))))))))))).))...))..))......-------- ( -14.20, z-score = -0.05, R) >droSec1.super_6 1411464 100 + 4358794 ACAUUUGGCCGAAUAUUCUUCAUUUUAGUAACAUUGUUAAUGCCUACUUAGAUAUAAACCACAUAUCAUACUGAAGUGAAGACGUAUACAAACAUACAUA-------- ................((((((((((((((.(((.....)))........(((((.......))))).)))))))))))))).((((......))))...-------- ( -19.40, z-score = -2.74, R) >droSim1.chr3R 1329175 108 + 27517382 ACAUUUGGCCGAAUAUUCUUCAUUUUAGUAACAUUGUUAAUGCCUACUUAGAUAUAAACCACAUAUCAUACUGAAGUGAAGACGUAUACAAACAUACAUAUGUAUGUA ..........(.((((((((((((((((((.(((.....)))........(((((.......))))).)))))))))))))).)))).)..((((((....)))))). ( -24.80, z-score = -3.19, R) >consensus ACAUUUGGCCGAAUAUUCUUCAUUUUAGUAACAUUGUUAAUGCCUACUUAGAUAUAAACCACAUAUCAUACUGAAGUGAAGACGUAUACAAACAUACAUA________ ................((((((((((((((.(((.....)))........(((((.......))))).)))))))))))))).((((......))))........... (-15.94 = -16.50 + 0.56)


| Location | 1,297,300 – 1,297,402 |
|---|---|
| Length | 102 |
| Sequences | 3 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 83.23 |
| Shannon entropy | 0.22540 |
| G+C content | 0.35371 |
| Mean single sequence MFE | -25.70 |
| Consensus MFE | -12.35 |
| Energy contribution | -15.47 |
| Covariance contribution | 3.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.79 |
| Structure conservation index | 0.48 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.20 |
| SVM RNA-class probability | 0.591474 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3R 1297300 102 + 27905053 CUGAAGUGAAGACGUAUACAAACACACAUA--------GCCACACACUGCCCAGUUAUUUUGAAAAAAUAUGGAAAAAUGUCAUAGACAUUUUUGGCAAAGCUGCCACAA .....(((.((.(((......)).......--------(((.........(((.....(((....)))..)))(((((((((...))))))))))))...)))..))).. ( -17.70, z-score = -0.41, R) >droSec1.super_6 1411534 102 + 4358794 CUGAAGUGAAGACGUAUACAAACAUACAUA--------GGUUCACAUUGCCCAAUUAUUUUGAAAAAAUAUGGAAAAAUGUCAUAGUGACAUUCGGGCAAGUUUUCACAG .....((((((((((((......))))...--------........((((((......(((....)))........((((((.....)))))).)))))))))))))).. ( -27.60, z-score = -3.90, R) >droSim1.chr3R 1329245 110 + 27517382 CUGAAGUGAAGACGUAUACAAACAUACAUAUGUAUGUAGGUUCACAUUGCCCAAUUAUUUUGAAAAAAUUUGGAAAAAUGGCAUAGUGACAUUCGGGCAAGUUUUCACAG .....((((((((((..((..((((((....))))))..))..)).((((((.....(((..((....))..))).((((.........)))).)))))))))))))).. ( -31.80, z-score = -4.04, R) >consensus CUGAAGUGAAGACGUAUACAAACAUACAUA________GGUUCACAUUGCCCAAUUAUUUUGAAAAAAUAUGGAAAAAUGUCAUAGUGACAUUCGGGCAAGUUUUCACAG .....((((((((((((......))))...................((((((.....((((.(.......).))))((((((.....)))))).)))))))))))))).. (-12.35 = -15.47 + 3.11)



Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:51:29 2011