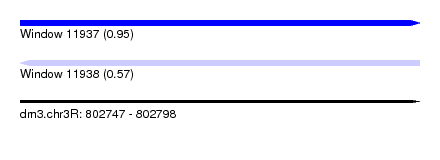

| Sequence ID | dm3.chr3R |
|---|---|
| Location | 802,747 – 802,798 |
| Length | 51 |
| Max. P | 0.948883 |

| Location | 802,747 – 802,798 |
|---|---|
| Length | 51 |
| Sequences | 6 |
| Columns | 61 |
| Reading direction | forward |
| Mean pairwise identity | 58.30 |
| Shannon entropy | 0.83912 |
| G+C content | 0.40813 |
| Mean single sequence MFE | -13.60 |
| Consensus MFE | -3.97 |
| Energy contribution | -4.80 |
| Covariance contribution | 0.84 |
| Combinations/Pair | 1.62 |
| Mean z-score | -1.84 |
| Structure conservation index | 0.29 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.54 |
| SVM RNA-class probability | 0.948883 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
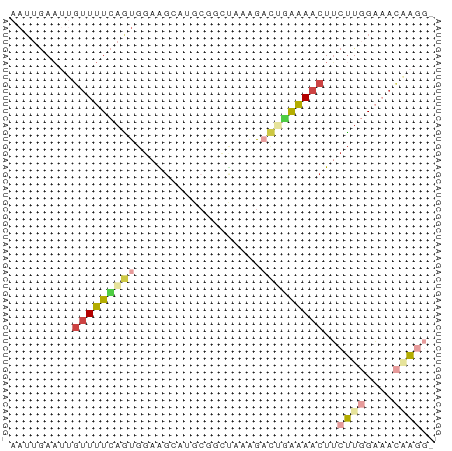
>dm3.chr3R 802747 51 + 27905053 AAUUGAAUUGUUUUCAGUGGAAGUG---------GGACUAAAAACUUCUUGGAAACCAGG- .........(((((((..((((((.---------.........)))))))))))))....- ( -10.50, z-score = -0.65, R) >droAna3.scaffold_13340 23019132 55 + 23697760 -----AAUUGUUUUUUCCAUAGCAAAGCUCACAUAGCCAAGGAUCUCCGUCGUCCUUCGU- -----..(((((........)))))..........((.((((((.......)))))).))- ( -7.40, z-score = -1.23, R) >droEre2.scaffold_4770 1097538 61 + 17746568 AAUUGAAUUGUUUUCAGUGGAGGCAUGCGGCUCAAGACGGAAAACUUCUUGGAAACAAUGG ......(((((((((((.((((...(.((........)).)...))))))))))))))).. ( -15.90, z-score = -2.07, R) >droSim1.chr3R_random 182595 60 + 1307089 AAUUGAAUUGUUUUCAGUGGAAGGAUGCGGCUCCGGACUGAAAACUUCUUGGAAACAAGG- .........(((((((((....(((......)))..)))))))))..((((....)))).- ( -19.20, z-score = -3.05, R) >droSec1.super_6 946911 60 + 4358794 AAUUGAAUUGUUUUCAGUGGAAGCAUGCGACUGCGGACUGAAAACUUCUUGGAAACAAGG- .........(((((((((....(((......)))..)))))))))..((((....)))).- ( -19.00, z-score = -3.65, R) >droGri2.scaffold_15074 3063202 55 - 7742996 AAUGAAUCCAUUUUGAAUGGAAG---GCAGGCAAAGUUGUGAAACUACUUAGUCAAGA--- ..((..(((((.....)))))..---.))(((.((((.(.....).)))).)))....--- ( -9.60, z-score = -0.39, R) >consensus AAUUGAAUUGUUUUCAGUGGAAGCAUGCGGCUAAAGACUGAAAACUUCUUGGAAACAAGG_ .........(((((((((..................)))))))))..((((....)))).. ( -3.97 = -4.80 + 0.84)

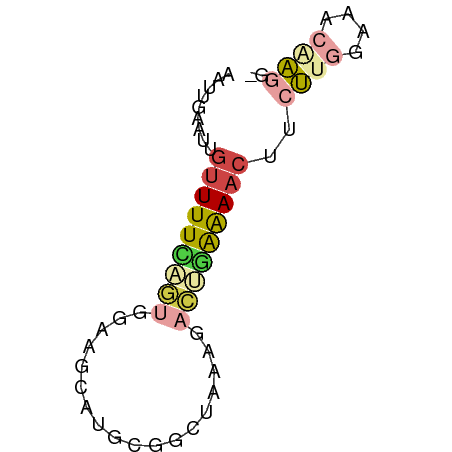
| Location | 802,747 – 802,798 |
|---|---|
| Length | 51 |
| Sequences | 6 |
| Columns | 61 |
| Reading direction | reverse |
| Mean pairwise identity | 58.30 |
| Shannon entropy | 0.83912 |
| G+C content | 0.40813 |
| Mean single sequence MFE | -9.58 |
| Consensus MFE | -2.49 |
| Energy contribution | -3.35 |
| Covariance contribution | 0.86 |
| Combinations/Pair | 1.56 |
| Mean z-score | -1.37 |
| Structure conservation index | 0.26 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.566328 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
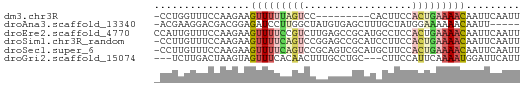
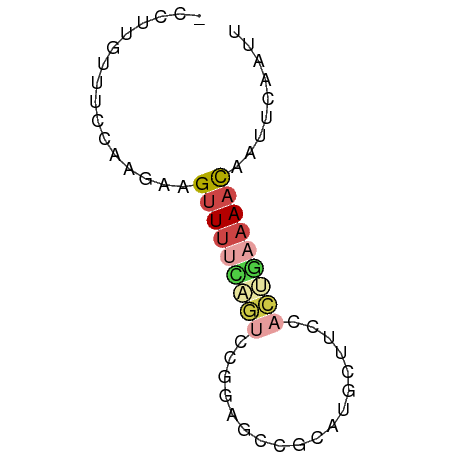
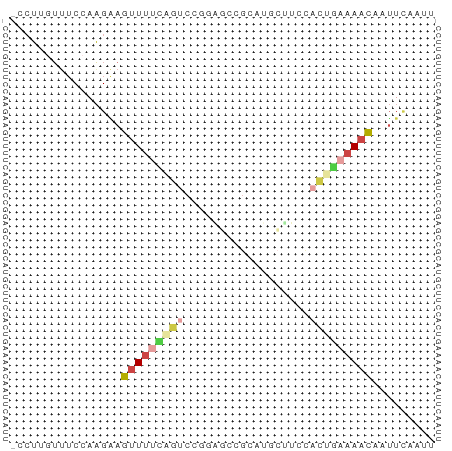
>dm3.chr3R 802747 51 - 27905053 -CCUGGUUUCCAAGAAGUUUUUAGUCC---------CACUUCCACUGAAAACAAUUCAAUU -..(((...)))....(((((((((..---------.......)))))))))......... ( -7.20, z-score = -0.77, R) >droAna3.scaffold_13340 23019132 55 - 23697760 -ACGAAGGACGACGGAGAUCCUUGGCUAUGUGAGCUUUGCUAUGGAAAAAACAAUU----- -.(....)..........(((.((((...(....)...)))).)))..........----- ( -7.80, z-score = 0.08, R) >droEre2.scaffold_4770 1097538 61 - 17746568 CCAUUGUUUCCAAGAAGUUUUCCGUCUUGAGCCGCAUGCCUCCACUGAAAACAAUUCAAUU ................((((((.((...(((........))).)).))))))......... ( -7.20, z-score = -0.27, R) >droSim1.chr3R_random 182595 60 - 1307089 -CCUUGUUUCCAAGAAGUUUUCAGUCCGGAGCCGCAUCCUUCCACUGAAAACAAUUCAAUU -.((((....))))..(((((((((..(((......)))....)))))))))......... ( -15.20, z-score = -3.65, R) >droSec1.super_6 946911 60 - 4358794 -CCUUGUUUCCAAGAAGUUUUCAGUCCGCAGUCGCAUGCUUCCACUGAAAACAAUUCAAUU -.((((....))))..(((((((((..(((......)))....)))))))))......... ( -15.00, z-score = -3.95, R) >droGri2.scaffold_15074 3063202 55 + 7742996 ---UCUUGACUAAGUAGUUUCACAACUUUGCCUGC---CUUCCAUUCAAAAUGGAUUCAUU ---...(((....((((...((......)).))))---..(((((.....))))).))).. ( -5.10, z-score = 0.36, R) >consensus _CCUUGUUUCCAAGAAGUUUUCAGUCCGGAGCCGCAUGCUUCCACUGAAAACAAUUCAAUU ................(((((((((..................)))))))))......... ( -2.49 = -3.35 + 0.86)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:50:25 2011