
| Sequence ID | dm3.chr3RHet |
|---|---|
| Location | 1,940,070 – 1,940,178 |
| Length | 108 |
| Max. P | 0.646992 |

| Location | 1,940,070 – 1,940,178 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 59.26 |
| Shannon entropy | 0.57362 |
| G+C content | 0.38067 |
| Mean single sequence MFE | -22.10 |
| Consensus MFE | -8.77 |
| Energy contribution | -10.00 |
| Covariance contribution | 1.23 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.40 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.33 |
| SVM RNA-class probability | 0.646992 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3RHet 1940070 108 + 2517507 GCGUGACAAUAAAAGGAUGUAACGGCGGAUCGUCGAG--UUCGGUAUGAAAGCUUGUGCUCCUUAUAAAAUAAGUAUUCAGAACAAACUGACUUAAAACGUAACAUAAAG ((((........(((((.(((..(((...((((((..--..))..))))..)))..))))))))......((((...((((......))))))))..))))......... ( -19.60, z-score = -0.14, R) >droSec1.super_61 148915 89 + 189864 GCGUGAUAAUAAGAAGAUA--ACGGCGAAUCGCCGGAGUUUCGAUUUAUGAGCUUGUGCUCUAGAUUGAAACAGUGGUCAAAG-AAAACUAA------------------ ...((((............--.(((((...)))))..(((((((((((.((((....)))))))))))))))....))))...-........------------------ ( -27.90, z-score = -3.68, R) >droYak2.chr2L_random 1873246 95 - 4064425 GCGUGAUAAUAAGAAGAUAU-ACGGCAAUUUGCCGGAGUUUCGAUU---------GUGCUCAUUAUUGAAUCAGUGGUCAAAGCAAAACUAAAACUUAUCUAACU----- ..............((((..-.((((.....))))((((((.....---------.((((.(((((((...)))))))...))))......))))))))))....----- ( -18.80, z-score = -0.83, R) >consensus GCGUGAUAAUAAGAAGAUAU_ACGGCGAAUCGCCGGAGUUUCGAUUU___AGCUUGUGCUCAUUAUUGAAUCAGUGGUCAAAGCAAAACUAA_____A__UAAC______ ...((((...............(((((...)))))....(((((((....(((....)))...)))))))......)))).............................. ( -8.77 = -10.00 + 1.23)

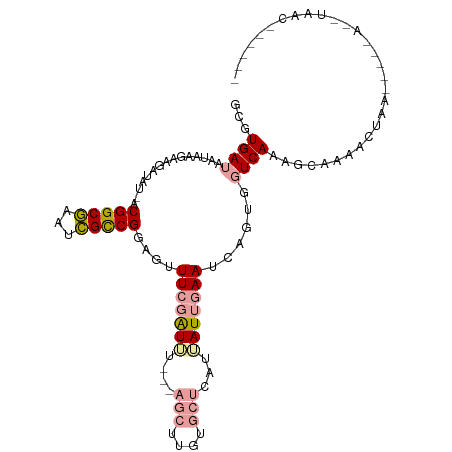

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:48:54 2011