| Sequence ID | dm3.chr3RHet |
|---|---|
| Location | 1,592,870 – 1,592,944 |
| Length | 74 |
| Max. P | 0.882799 |

| Location | 1,592,870 – 1,592,944 |
|---|---|
| Length | 74 |
| Sequences | 4 |
| Columns | 82 |
| Reading direction | reverse |
| Mean pairwise identity | 55.01 |
| Shannon entropy | 0.70017 |
| G+C content | 0.58381 |
| Mean single sequence MFE | -24.54 |
| Consensus MFE | -12.46 |
| Energy contribution | -12.03 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.69 |
| Mean z-score | -0.82 |
| Structure conservation index | 0.51 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.06 |
| SVM RNA-class probability | 0.882799 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
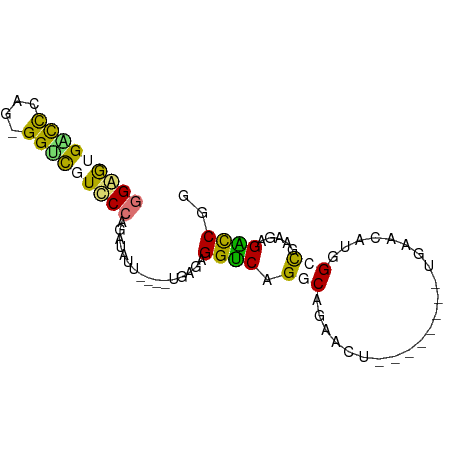
>dm3.chr3RHet 1592870 74 - 2517507 GGAAUGGUUGAGUGGCCUUUCAGGCU-------UGAGAGGCCGGUCAGGUCUACC-CAAGUGAGGAAGGUUAGAGAGGCCGG ....(((((...(((((((((..(((-------((.((((((.....))))).).-)))))..)))))))))....))))). ( -28.90, z-score = -2.29, R) >droYak2.chr2L_random 3129081 81 - 4064425 GGAGAGACCGUCCGGGUCUACCAGUUAACAGGGUACGAGGUCGGUCAGGCCUACCACAGAUGAGAACAGUU-UAGAGGCCGG .....(((((.((..((...((........))..))..)).))))).(((((........((....))...-...))))).. ( -24.24, z-score = -0.34, R) >droSec1.super_62 53668 69 - 174595 GGGGUGACCCAG-GGUCGUCCCAGAUAUU----UGAUGGGUCAGGCAGAACU--------UGAACGUGGCCGAGGAGAUCGU (((....)))..-(((((.((((......----...)))).).(((...((.--------.....)).))).....)))).. ( -24.10, z-score = -0.97, R) >droSim1.chr2h_random 2095484 69 - 3178526 GGGGUGGCCCAG-GGUCGUCCCAGAUAUU----UGAUGGGUCAGGCAGAACU--------UGAACGUGGCCGAGGAGAUCGU (((..((((...-))))..))).(((...----...(.(((((.(.......--------....).))))).)....))).. ( -20.90, z-score = 0.31, R) >consensus GGAGUGACCCAG_GGUCGUCCCAGAUAUU____UGAGAGGUCAGGCAGAACU________UGAACAUGGCCGAAGAGACCGG (((((((((....)))))))))................((((.(((......................))).....)))).. (-12.46 = -12.03 + -0.44)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:48:37 2011