
| Sequence ID | dm3.chr3RHet |
|---|---|
| Location | 749,776 – 749,836 |
| Length | 60 |
| Max. P | 0.694254 |

| Location | 749,776 – 749,836 |
|---|---|
| Length | 60 |
| Sequences | 3 |
| Columns | 70 |
| Reading direction | forward |
| Mean pairwise identity | 71.63 |
| Shannon entropy | 0.40182 |
| G+C content | 0.32386 |
| Mean single sequence MFE | -9.90 |
| Consensus MFE | -6.68 |
| Energy contribution | -6.47 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.23 |
| Mean z-score | -1.30 |
| Structure conservation index | 0.68 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.43 |
| SVM RNA-class probability | 0.694254 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
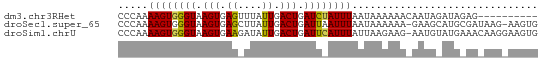
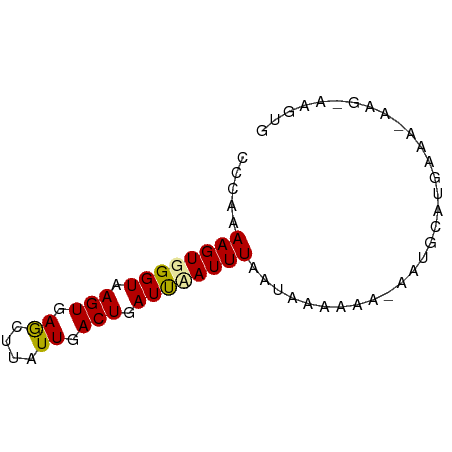
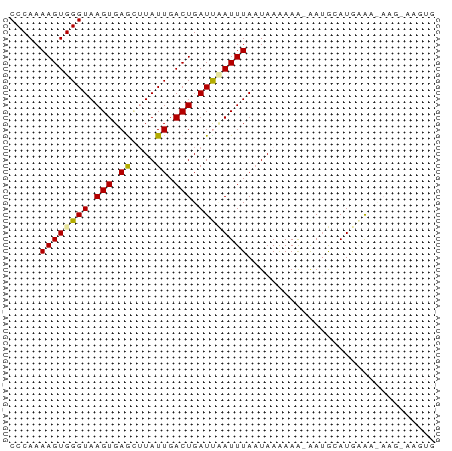
>dm3.chr3RHet 749776 60 + 2517507 CCCAAAAGUGGGUAAGUGAGUUUAUUGACUGAUCUAUUUAAUAAAAAACAAUAGAUAGAG---------- ((((....))))....(.((((....)))).)((((((((...........)))))))).---------- ( -9.90, z-score = -1.68, R) >droSec1.super_65 147185 68 - 171001 CCCAAAAGUGGGUAAGUGAGCUUAUUGACUGAUUAAUUUAAUAAAAAA-GAAGCAUGCGAUAAG-AAGUG ((((....))))........(((((((..((.....(((......)))-....))..)))))))-..... ( -9.60, z-score = -1.08, R) >droSim1.chrU 13436858 69 - 15797150 CCCAAAAGUGGGUAAGUGAAGAUAUUGACUGAUUCAUUUAUUAAGAAG-AAUGUAUGAAACAAGGAAGUG ((((....))))..........((((..((..(((((.((((......-)))).)))))...))..)))) ( -10.20, z-score = -1.15, R) >consensus CCCAAAAGUGGGUAAGUGAGCUUAUUGACUGAUUAAUUUAAUAAAAAA_AAUGCAUGAAA_AAG_AAGUG .....((((((((.(((.((....)).))).))))))))............................... ( -6.68 = -6.47 + -0.22)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:48:00 2011