| Sequence ID | dm3.chr3RHet |
|---|---|
| Location | 734,515 – 734,572 |
| Length | 57 |
| Max. P | 0.814916 |

| Location | 734,515 – 734,572 |
|---|---|
| Length | 57 |
| Sequences | 6 |
| Columns | 57 |
| Reading direction | reverse |
| Mean pairwise identity | 58.83 |
| Shannon entropy | 0.82640 |
| G+C content | 0.47722 |
| Mean single sequence MFE | -15.48 |
| Consensus MFE | -3.80 |
| Energy contribution | -4.66 |
| Covariance contribution | 0.86 |
| Combinations/Pair | 1.82 |
| Mean z-score | -1.76 |
| Structure conservation index | 0.25 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.78 |
| SVM RNA-class probability | 0.814916 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3RHet 734515 57 - 2517507 UACAUAGAGUUAGCCUCUGUAAGUCCCAUCGCGUAUUCUGCAGAGGACUGCUCAAAG ......((((...(((((((((((.(......).))).))))))))...)))).... ( -17.00, z-score = -2.64, R) >anoGam1.chr2R 44536208 57 + 62725911 GGCAAUAAAACAGCGGCCGGAAUUCGUUCCACACCAAUUUUGCAGCAAUGCUCGAAG (((............)))((((....))))...........(((....)))...... ( -10.40, z-score = -0.03, R) >dp4.chr4_group4 6136433 53 - 6586962 CACACGGACUUUUUUGAAAAAAAUAAAAUUUCGUAAGAAA-ACCGAUCUCUUUG--- ....(((....((((....)))).....((((....))))-.))).........--- ( -8.00, z-score = -1.47, R) >droEre2.scaffold_4929 23987585 57 - 26641161 UACAUCGAGCUAGCUUCUGCAAGUCCCAUCGCGUGUUCUGCAGAGGGCUGCUCAAAG ......((((.((((((((((.(..((.....).)..))))))).)))))))).... ( -19.10, z-score = -1.69, R) >droYak2.chrU 17479659 57 - 28119190 UACAUAGAGUCAGCUCCUGCAAGUCCCAUCGCGUGUUCUGCAGAGGGCUGCUCUAAG ....(((((.(((((((((((.(..((.....).)..)))))).))))))))))).. ( -22.90, z-score = -3.29, R) >droSim1.chrU 13094116 57 + 15797150 UACAUAGAGUUAGCCCUUGCAAGUCCCAUCGCGUGUUCUGCAGAGGGAUGCUCAAAG ......((((...((((((((.(..((.....).)..))))).))))..)))).... ( -15.50, z-score = -1.44, R) >consensus UACAUAGAGUUAGCUCCUGCAAGUCCCAUCGCGUGUUCUGCAGAGGGCUGCUCAAAG ......((((...((.(((((.................))))).))...)))).... ( -3.80 = -4.66 + 0.86)

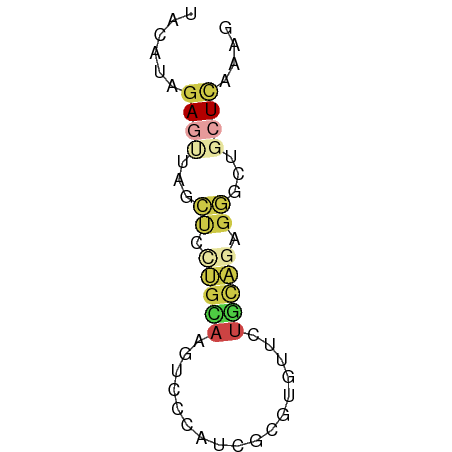

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:47:59 2011