
| Sequence ID | dm3.chr3RHet |
|---|---|
| Location | 545,806 – 545,898 |
| Length | 92 |
| Max. P | 0.652414 |

| Location | 545,806 – 545,898 |
|---|---|
| Length | 92 |
| Sequences | 3 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 59.62 |
| Shannon entropy | 0.53369 |
| G+C content | 0.50141 |
| Mean single sequence MFE | -24.10 |
| Consensus MFE | -9.84 |
| Energy contribution | -12.07 |
| Covariance contribution | 2.23 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.41 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.34 |
| SVM RNA-class probability | 0.652414 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3RHet 545806 92 + 2517507 UUGGCAAAGC--UUGAAUUCUCUGCAACUCGUGAUUCCAUGUUGUCCCCCACAGAUGGAACAACCUCCAUGCCGAUCAUCCUGUUGAUCGCAUU ..((((..((--..((....)).)).....((..((((((.((((.....))))))))))..)).....))))(((((......)))))..... ( -22.10, z-score = -1.41, R) >droSim1.chrU 10350766 77 + 15797150 GCAGAAGAGUGGUUAAAUUCUUUGCAGUCCUUUAUCCAGUGUUGUCCCUCACACAUGGGGCAACGUCUAGGGCGGUC----------------- (((((.((((......))))))))).(((((......(((((((((((........))))))))).)))))))....----------------- ( -28.10, z-score = -3.18, R) >droAna3.scaffold_12947 1154382 77 - 1574637 GUAGAGACGCUGUUGAAUUCUUUGCAGUCCCUGAUCUGGUGUUGUCCCUCGCAUAUAGGGCAAAAUCUGGGGCGGUC----------------- (((((((..(....)...))))))).(((((.(((......(((.((((.......))))))).))).)))))....----------------- ( -22.10, z-score = -0.48, R) >consensus GUAGAAAAGC_GUUGAAUUCUUUGCAGUCCCUGAUCCAGUGUUGUCCCUCACACAUGGGGCAACAUCUAGGGCGGUC_________________ ..........................(((((.(((.....((((((((........))))))))))).)))))..................... ( -9.84 = -12.07 + 2.23)

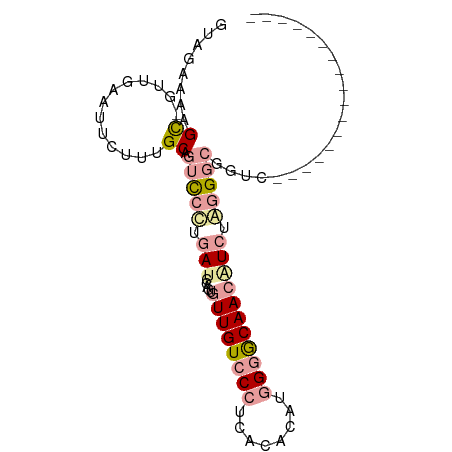

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:47:42 2011