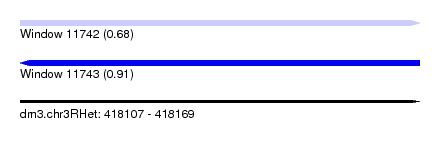

| Sequence ID | dm3.chr3RHet |
|---|---|
| Location | 418,107 – 418,169 |
| Length | 62 |
| Max. P | 0.912640 |

| Location | 418,107 – 418,169 |
|---|---|
| Length | 62 |
| Sequences | 4 |
| Columns | 62 |
| Reading direction | forward |
| Mean pairwise identity | 70.70 |
| Shannon entropy | 0.49055 |
| G+C content | 0.71668 |
| Mean single sequence MFE | -23.48 |
| Consensus MFE | -10.61 |
| Energy contribution | -11.30 |
| Covariance contribution | 0.69 |
| Combinations/Pair | 1.25 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.40 |
| SVM RNA-class probability | 0.680227 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
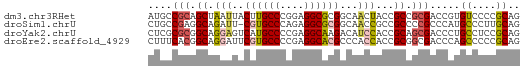
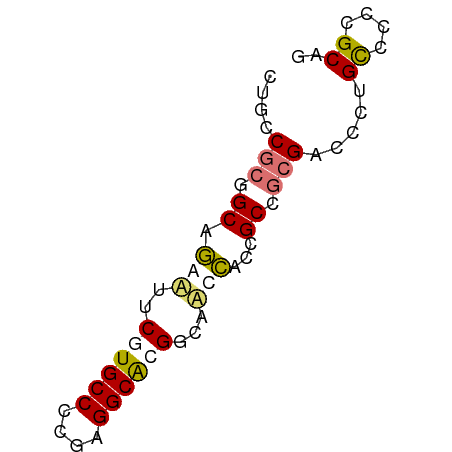
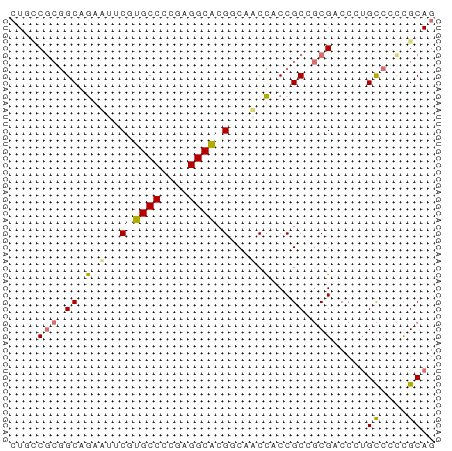
>dm3.chr3RHet 418107 62 + 2517507 AUGCCGCAGCUAAUUACUUGCCCGGAGGCGCGGCAACUACCGCCGCGACCGUGUCCCCGCAG .(((.((((........))))..(((((((((((.......)))))).))...)))..))). ( -20.90, z-score = -0.50, R) >droSim1.chrU 939901 61 - 15797150 CUGCCGAGGCAGAUU-CGUGCCCAGAGGCGCGGCAACCGCCGCCCCGCCCAUGCCCUUGCAG ((((...((((....-((.(((....)))(((((....)))))..))....))))...)))) ( -24.50, z-score = -1.69, R) >droYak2.chrU 6215484 62 + 28119190 CUCGCGCGGCAGGAGUCAUGCCCCGAGGCAAGACAUCCACCGCAGCGACCCUGCCUCCGCAG .((((((((..((((((.((((....)))).))).))).)))).))))..((((....)))) ( -28.80, z-score = -3.67, R) >droEre2.scaffold_4929 967677 62 - 26641161 CUUUCACGGCAGGAUUCGUGCCCCGAGGCACGCCCACCACCGCGGCGACCCAGCCCCCGCAG .......((..((...((((((....))))))))..))...((((.(........))))).. ( -19.70, z-score = -1.04, R) >consensus CUGCCGCGGCAGAAUUCGUGCCCCGAGGCACGGCAACCACCGCCGCGACCCUGCCCCCGCAG ....(((.((.(((..((((((....))))))...)))...)).))).....((....)).. (-10.61 = -11.30 + 0.69)
| Location | 418,107 – 418,169 |
|---|---|
| Length | 62 |
| Sequences | 4 |
| Columns | 62 |
| Reading direction | reverse |
| Mean pairwise identity | 70.70 |
| Shannon entropy | 0.49055 |
| G+C content | 0.71668 |
| Mean single sequence MFE | -31.10 |
| Consensus MFE | -16.80 |
| Energy contribution | -18.30 |
| Covariance contribution | 1.50 |
| Combinations/Pair | 1.28 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.54 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.23 |
| SVM RNA-class probability | 0.912640 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
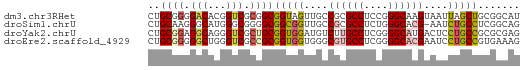
>dm3.chr3RHet 418107 62 - 2517507 CUGCGGGGACACGGUCGCGGCGGUAGUUGCCGCGCCUCCGGGCAAGUAAUUAGCUGCGGCAU .((.(....).))((((((((..((.(((((.((....))))))).))....)))))))).. ( -27.30, z-score = -0.87, R) >droSim1.chrU 939901 61 + 15797150 CUGCAAGGGCAUGGGCGGGGCGGCGGUUGCCGCGCCUCUGGGCACG-AAUCUGCCUCGGCAG ((((..(((((.(..((..(((((....)))))(((....))).))-..).)))))..)))) ( -30.90, z-score = -1.49, R) >droYak2.chrU 6215484 62 - 28119190 CUGCGGAGGCAGGGUCGCUGCGGUGGAUGUCUUGCCUCGGGGCAUGACUCCUGCCGCGCGAG ((((....))))..((((.((((((((.(((.((((....)))).)))))).))))))))). ( -33.90, z-score = -2.64, R) >droEre2.scaffold_4929 967677 62 + 26641161 CUGCGGGGGCUGGGUCGCCGCGGUGGUGGGCGUGCCUCGGGGCACGAAUCCUGCCGUGAAAG ((((((.(((...))).)))))).((..((((((((....))))))...))..))....... ( -32.30, z-score = -2.00, R) >consensus CUGCGGGGGCAGGGUCGCGGCGGUGGUUGCCGCGCCUCGGGGCACGAAACCUGCCGCGGCAG ..((((.(((...))).))))(((((....((((((....))))))....)))))....... (-16.80 = -18.30 + 1.50)
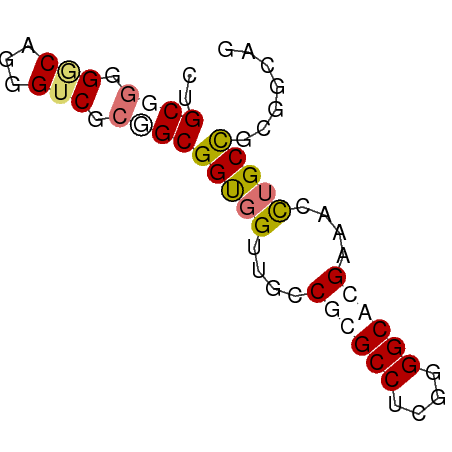
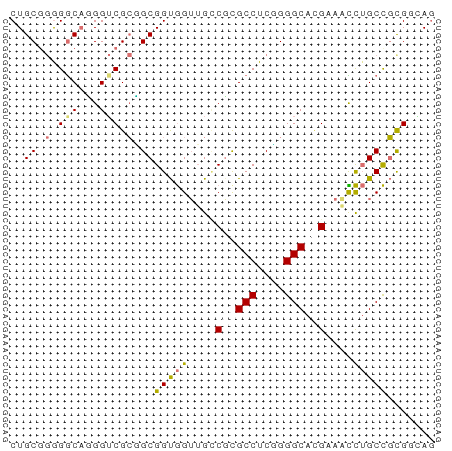
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:47:40 2011