| Sequence ID | dm3.chr3L |
|---|---|
| Location | 24,479,092 – 24,479,158 |
| Length | 66 |
| Max. P | 0.978240 |

| Location | 24,479,092 – 24,479,158 |
|---|---|
| Length | 66 |
| Sequences | 4 |
| Columns | 68 |
| Reading direction | forward |
| Mean pairwise identity | 52.69 |
| Shannon entropy | 0.72016 |
| G+C content | 0.59747 |
| Mean single sequence MFE | -28.93 |
| Consensus MFE | -8.95 |
| Energy contribution | -10.33 |
| Covariance contribution | 1.38 |
| Combinations/Pair | 1.60 |
| Mean z-score | -2.32 |
| Structure conservation index | 0.31 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.99 |
| SVM RNA-class probability | 0.978240 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
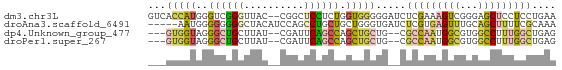
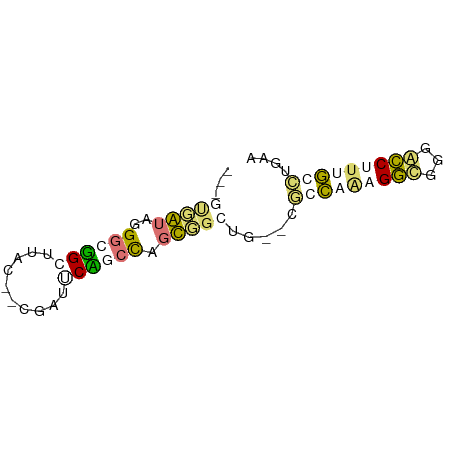
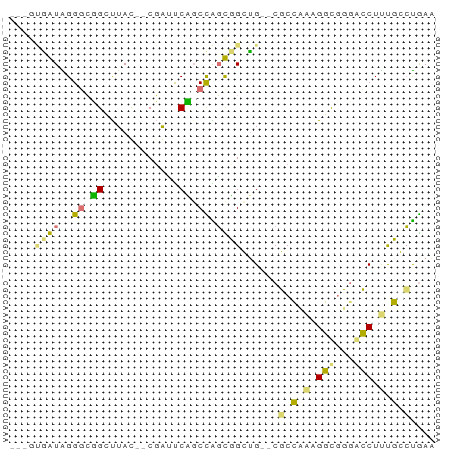
>dm3.chr3L 24479092 66 + 24543557 GUCACCAUGGGUCGGGUUAC--CGGCUCCUCUGGUGGGGGAUCUCGAAAGUCGGGAGCUCCUCCUGAA ..(((((.(((((((....)--)))).))..)))))((((.(((((.....))))).))))....... ( -29.80, z-score = -1.62, R) >droAna3.scaffold_6491 3087 63 + 3293 -----AAUGGGGGGGGCUACAUCCAGCCUGCUGCUGGGUGAUCUCGUGAGUUUGCAGCUUUUCGCAAA -----..((.(..(((((.((((((((.....))))))))....(....).....)))))..).)).. ( -24.90, z-score = -1.76, R) >dp4.Unknown_group_477 5753 61 + 9680 ---GUGGUAGGGCUGCUUAU--CGAUUCAGCCAGCUGCUG--CGCCAAUGGCGUGGCCUUUGGCUGAG ---.(((((((....)))))--))..((((((((..((..--((((...))))..))..)))))))). ( -30.50, z-score = -2.94, R) >droPer1.super_267 5384 61 - 27109 ---GUGGUAGGGCUGCUUAU--CGAUUCAGCCAGCUGCUG--CGCCAAUGGCGUGGCCUUUGGCUGAG ---.(((((((....)))))--))..((((((((..((..--((((...))))..))..)))))))). ( -30.50, z-score = -2.94, R) >consensus ___GUGAUAGGGCGGCUUAC__CGAUUCAGCCAGCGGCUG__CGCCAAAGGCGGGACCUUUGCCUGAA ...(((((..((((((..........)))))).))))).....(((((((((...))))))))).... ( -8.95 = -10.33 + 1.38)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:47:18 2011