
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 24,169,892 – 24,169,955 |
| Length | 63 |
| Max. P | 0.554578 |

| Location | 24,169,892 – 24,169,955 |
|---|---|
| Length | 63 |
| Sequences | 5 |
| Columns | 64 |
| Reading direction | reverse |
| Mean pairwise identity | 62.15 |
| Shannon entropy | 0.67870 |
| G+C content | 0.41679 |
| Mean single sequence MFE | -14.24 |
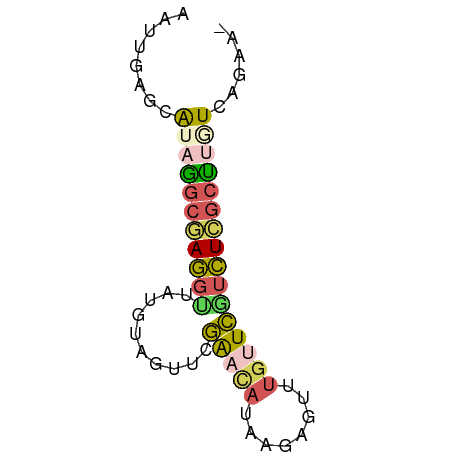
| Consensus MFE | -5.82 |
| Energy contribution | -6.38 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.69 |
| Mean z-score | -0.95 |
| Structure conservation index | 0.41 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.12 |
| SVM RNA-class probability | 0.554578 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
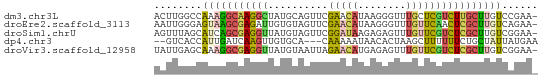
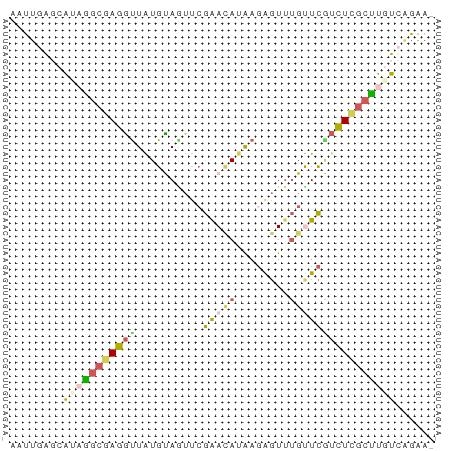
>dm3.chr3L 24169892 63 - 24543557 ACUUGGCCAAAGGCAAGGCUAUGCAGUUCGAACAUAAGGGUUUGCUCGUCUUGCUUGUCCGAA- ..((((....(((((((((...((((..(........)..).)))..)))))))))..)))).- ( -19.40, z-score = -1.18, R) >droEre2.scaffold_3113 2845 63 + 9025 AAUUGGGAGUAAGCGAGAUUGUGUAGUUCGAACAUAAGGGUUUGUUCAACUCGCUUGUCAGAA- ......((.((((((((.(((...((..(........)..))....)))))))))))))....- ( -15.70, z-score = -1.69, R) >droSim1.chrU 10613725 63 + 15797150 AGUUUAGCAUCAGCGAGGUUAUGUAGUUCGGAUAAGAGAGUUUGUUCGUCUCGCUUGUCGGAA- ..((..(((..(((((((..........((((((((....))))))))))))))))))..)).- ( -15.40, z-score = -1.20, R) >dp4.chr3 87183 59 - 19779522 --GUCACCAUUGAUCAAGUUGUGCA---CAAAAAUAACACUAAGCUUUUUUCUGCUAUUAUGAA --...........(((.(((((...---.....)))))....(((........)))....))). ( -6.70, z-score = 0.11, R) >droVir3.scaffold_12958 1550939 63 - 3547706 UAUUGAGCAAAGGCGAGGUUAUGUAAUUAGAACAUGAGAGUUUGUUCGUCUCGCUUGUCGGAA- ..((((....((((((((.((......))(((((........))))).)))))))).))))..- ( -14.00, z-score = -0.79, R) >consensus AAUUGAGCAUAGGCGAGGUUAUGUAGUUCGAACAUAAGAGUUUGUUCGUCUCGCUUGUCAGAA_ ........(((((((((((..........(((((........))))))))))))))))...... ( -5.82 = -6.38 + 0.56)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:46:58 2011