| Sequence ID | dm3.chr2L |
|---|---|
| Location | 6,186,351 – 6,186,459 |
| Length | 108 |
| Max. P | 0.918719 |

| Location | 6,186,351 – 6,186,459 |
|---|---|
| Length | 108 |
| Sequences | 11 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 72.69 |
| Shannon entropy | 0.58212 |
| G+C content | 0.55261 |
| Mean single sequence MFE | -21.95 |
| Consensus MFE | -9.17 |
| Energy contribution | -9.58 |
| Covariance contribution | 0.41 |
| Combinations/Pair | 1.24 |
| Mean z-score | -1.87 |
| Structure conservation index | 0.42 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.27 |
| SVM RNA-class probability | 0.918719 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
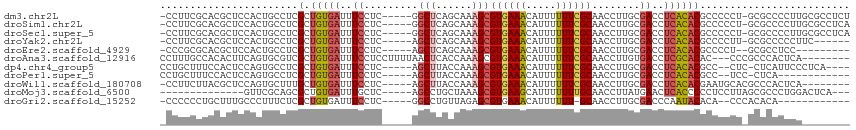
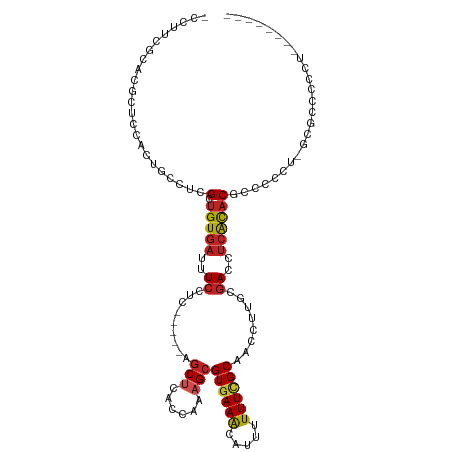
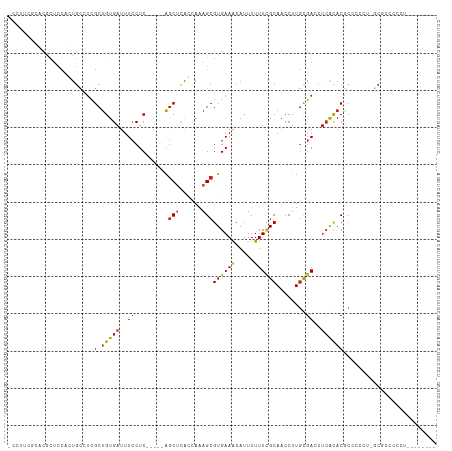
>dm3.chr2L 6186351 108 + 23011544 -CCUUCGCACGCUCCACUGCCUCGCUGUGAUUUCCUC-----GGCUCAGCAAAGCGUGAAACAUUUUUUCGCAACCUUGCGACCUCACACGCCCCCU-GCGCCCCUUGCGCCUCU -......((((((....(((...((((.(.....).)-----)))...))).))))))..........(((((....)))))...............-((((.....)))).... ( -25.80, z-score = -1.59, R) >droSim1.chr2L 5979178 108 + 22036055 -CCUUCGCACGCUCCACUGCCUCGCUGUGAUUUCCUC-----GGCUCAGCAAAGCGUGAAACAUUUUUUCGCAACCUUGCGACCUCACACGCCCCCU-GCGCCCCUUGCGCCUCA -......((((((....(((...((((.(.....).)-----)))...))).))))))..........(((((....)))))...............-((((.....)))).... ( -25.80, z-score = -1.52, R) >droSec1.super_5 4257589 108 + 5866729 -CCUUCGCACGCUCCACUGCCUCGCUGUGAUUUCCUC-----GGCUCAGCAAAGCGUGAAACAUUUUUUCGCAACCUUGCGACCUCACACGCCCCCU-GCGCCCCUUGCGCCUCA -......((((((....(((...((((.(.....).)-----)))...))).))))))..........(((((....)))))...............-((((.....)))).... ( -25.80, z-score = -1.52, R) >droYak2.chr2L 15601073 102 - 22324452 -CCUUCGCACGCUCCACUGCCUCGCUGUGAUUUCCUC-----AGCUCAGCAAAGCGUGAAACAUUUUUUCGCAACCUUGCGACCUCACACGCCCCUU-GCGCCCCCUUC------ -......((((((....(((...((((.(.....).)-----)))...))).))))))...........(((((....(((........)))...))-)))........------ ( -23.30, z-score = -2.48, R) >droEre2.scaffold_4929 15113999 98 + 26641161 -CCCGCGCACGCUCCACUGCCUCGCUGUGAUUUCCUC-----AGCUCAGCAAAGCGUGAAACAUUUUUUCGCAACCUUGCGACCUCACACGCCCCU--GCGCCUCC--------- -...(((((((((....(((...((((.(.....).)-----)))...))).)))))...........(((((....)))))..............--))))....--------- ( -25.80, z-score = -2.53, R) >droAna3.scaffold_12916 8163491 104 - 16180835 CCUUUGCCACACUUCAGUGCGUCGCUGUGAUUUCCUCCUUUUAACUCACCAAAGCGUGAAACAUUUUUUCGCAACCUUGUGACCUCGCACAC---CCCGCCCACUCA-------- ................(((((((((.((((...............))))......((((((.....))))))......)))))...))))..---............-------- ( -17.06, z-score = -1.32, R) >dp4.chr4_group5 49135 103 - 2436548 CCUGCUUUCCACUCCAGUGCCUCGCUGUGAUUUCCUC-----AGCUUACCAAAGCGUGAAACAUUUUUUCGCAACCUUGCGACCUCACACGCC--CUC-CUCAUUCCCUCA---- .........(((....)))....(((((((.....((-----(((((....)))).))).........(((((....)))))..))))).)).--...-............---- ( -18.00, z-score = -2.46, R) >droPer1.super_5 50073 95 - 6813705 CCUGCUUUCCACUCCAGUGCCUCGCUGUGAUUUCCUC-----AGCUUACCAAAGCGUGAAACAUUUUUUCGCAACCUUGCGACCUCACACGCC--UCC-CUCA------------ .........(((....)))....(((((((.....((-----(((((....)))).))).........(((((....)))))..))))).)).--...-....------------ ( -18.00, z-score = -2.14, R) >droWil1.scaffold_180708 5026080 101 - 12563649 -CCUUCUUACGCUCCAGUGCUUUGCUGUGAUUUCCUC-----AGCUUACCAAAGCGUGAAACAUUUUUUCGCAACCUUGCGACCUCACACGAAUGCACGCCCACUCA-------- -...............((((((((.(((((.....((-----(((((....)))).))).........(((((....)))))..))))))))).)))).........-------- ( -24.00, z-score = -3.06, R) >droMoj3.scaffold_6500 18982744 93 - 32352404 --------------GUUCGCAGCGCUGUGAUUUGCUC-----AGCCUGCUAAAGCGUGAAGCAUUUUUUUGCAACCUUAUGAACUCACCCCCUCCUUAGCGCCCUGGACUCA--- --------------((((((((.((((.(.....).)-----)))))))....((((...(((......))).......((....))...........))))...))))...--- ( -22.70, z-score = -0.98, R) >droGri2.scaffold_15252 14535738 94 - 17193109 -CCCCCCUGCUUUGCCCUUUCUCGCUGUGAUUUCCUC-----GGCCUGUUAGAGCGUGAAACAUUUUUU-GCAACCUUGCGACCCAAUACACA--CCCACACA------------ -......((..(((...((((.((((.((((..(...-----.)...)))).)))).))))......((-(((....)))))..)))..))..--........------------ ( -15.20, z-score = -1.00, R) >consensus _CCUUCGCACGCUCCACUGCCUCGCUGUGAUUUCCUC_____AGCUCACCAAAGCGUGAAACAUUUUUUCGCAACCUUGCGACCUCACACGCCCCCU_GCGCCCCCU________ .......................(.(((((..((.........(((......)))((((((.....))))))........))..))))))......................... ( -9.17 = -9.58 + 0.41)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 21:20:38 2011