| Sequence ID | dm3.chr3L |
|---|---|
| Location | 23,243,754 – 23,243,809 |
| Length | 55 |
| Max. P | 0.909849 |

| Location | 23,243,754 – 23,243,809 |
|---|---|
| Length | 55 |
| Sequences | 5 |
| Columns | 68 |
| Reading direction | forward |
| Mean pairwise identity | 50.00 |
| Shannon entropy | 0.81801 |
| G+C content | 0.47309 |
| Mean single sequence MFE | -16.26 |
| Consensus MFE | -4.06 |
| Energy contribution | -4.34 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.75 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.25 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.21 |
| SVM RNA-class probability | 0.909849 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
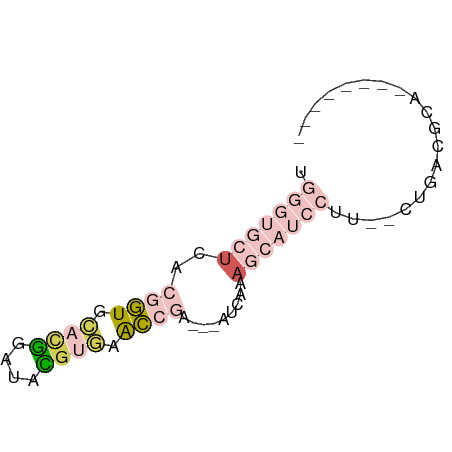
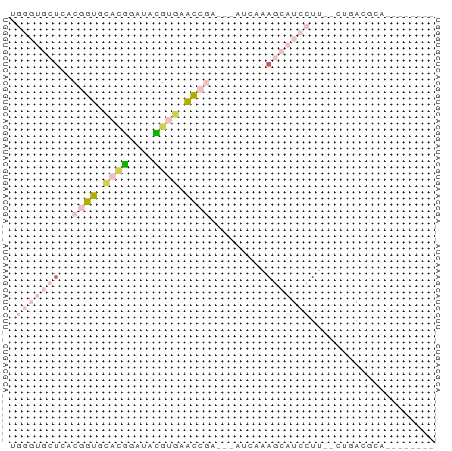
>dm3.chr3L 23243754 55 + 24543557 UGGGUGCUGACGGUGCACGGAUACGUGAACCAA---AUAUAAGCAUCCUU--CUGACGCC-------- .(((((((...(((.((((....)))).)))..---.....)))))))..--........-------- ( -18.80, z-score = -2.62, R) >droSim1.chr3L 22414225 55 + 22553184 UGGGUGCUCACGGUGCACGGAUACGUGAACCGA---ACCAAAGCAUCCUU--CUGACGCA-------- .(((((((..((((.((((....)))).)))).---.....)))))))..--........-------- ( -20.30, z-score = -2.42, R) >droSec1.super_41 160880 55 + 296808 UGGGUGCUCACGGUGCACGGAUACGUGAACCGA---ACCAAAGCAUCCUU--CUGACGCA-------- .(((((((..((((.((((....)))).)))).---.....)))))))..--........-------- ( -20.30, z-score = -2.42, R) >droPer1.super_43 36321 53 + 623856 ---------UCUGUGCCUAGAAUUGAAGGCUGAGAGAUCUGAAUGUCUUUUGCUCAGCCUUC------ ---------((((....))))...((((((((((((((......))))....))))))))))------ ( -17.70, z-score = -2.50, R) >apiMel3.Group15 4796306 62 - 7856270 UCAACCAUUCCAUUAUCUGAAAAUUUAGGAC-A---AUCAGAACAAAAUU--GUAAUUCUAUAAUACA ...............(((((...........-.---.))))).....(((--(((....))))))... ( -4.22, z-score = 0.03, R) >consensus UGGGUGCUCACGGUGCACGGAUACGUGAACCGA___AUCAAAGCAUCCUU__CUGACGCA________ ..........((((.((((....)))).)))).................................... ( -4.06 = -4.34 + 0.28)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:46:16 2011