
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 22,751,922 – 22,751,991 |
| Length | 69 |
| Max. P | 0.791449 |

| Location | 22,751,922 – 22,751,991 |
|---|---|
| Length | 69 |
| Sequences | 5 |
| Columns | 70 |
| Reading direction | forward |
| Mean pairwise identity | 67.16 |
| Shannon entropy | 0.55757 |
| G+C content | 0.42462 |
| Mean single sequence MFE | -14.14 |
| Consensus MFE | -5.09 |
| Energy contribution | -5.29 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.82 |
| Structure conservation index | 0.36 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.626059 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
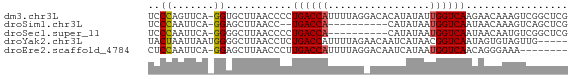
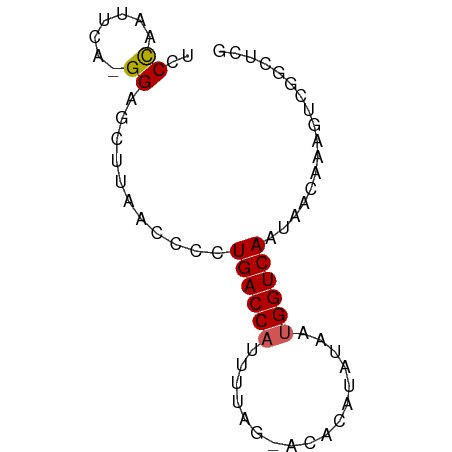
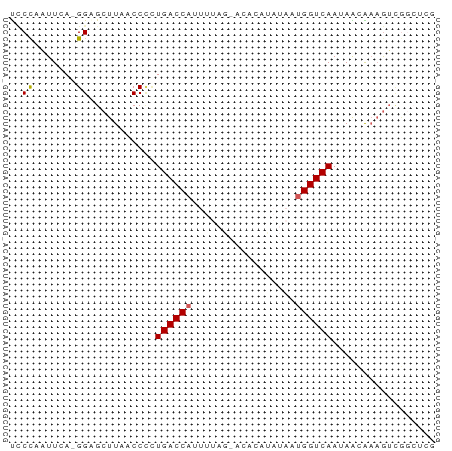
>dm3.chr3L 22751922 69 + 24543557 UCCCAGUUCA-GGUGCUUAACCCCUGACCAUUUUAGGACACAUAUAUUGGUCAAGAACAAAGUCGGCUCG .....((((.-(((.....)))..((((((...((.......))...)))))).))))............ ( -14.00, z-score = -0.65, R) >droSim1.chr3L 22040030 57 + 22553184 UCCCAAUUCA-GGAGCUUAACC--UGACCA----------CAUAUAAUGGUCAAUAACAAAGUCAGCUCG ..........-.(((((.....--((((((----------.......))))))...........))))). ( -10.59, z-score = -2.17, R) >droSec1.super_11 2672551 59 + 2888827 UCCCAAUUCA-GGGGCUUAACCCCUGACCA----------CAUAUAAUGGUCAAUAACAAUGUCGGCUCG ..((.....(-((((.....)))))(((((----------.......)))))............)).... ( -15.30, z-score = -2.09, R) >droYak2.chr3L 23243090 65 + 24197627 UACUAAUUAAUGGGGCUUAACCUCUGACCAUUUUAGAACAAUCAUAACGGUCAAUAGUGUAGUUG----- (((((......((((.....))))(((((.((...((....))..)).))))).)))))......----- ( -11.30, z-score = -0.65, R) >droEre2.scaffold_4784 22403994 61 + 25762168 CUCCAAUUCA-GGAGCUUAACCCUUGACCAUUUUAGGACAAUCAUAAUGGUCAACAGGGAAA-------- ((((......-)))).....((((((((((((....(.....)..))))))))..))))...-------- ( -19.50, z-score = -3.52, R) >consensus UCCCAAUUCA_GGAGCUUAACCCCUGACCAUUUUAG_ACACAUAUAAUGGUCAAUAACAAAGUCGGCUCG ........................((((((.................))))))................. ( -5.09 = -5.29 + 0.20)
| Location | 22,751,922 – 22,751,991 |
|---|---|
| Length | 69 |
| Sequences | 5 |
| Columns | 70 |
| Reading direction | reverse |
| Mean pairwise identity | 67.16 |
| Shannon entropy | 0.55757 |
| G+C content | 0.42462 |
| Mean single sequence MFE | -17.03 |
| Consensus MFE | -6.61 |
| Energy contribution | -6.53 |
| Covariance contribution | -0.08 |
| Combinations/Pair | 1.33 |
| Mean z-score | -1.86 |
| Structure conservation index | 0.39 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.70 |
| SVM RNA-class probability | 0.791449 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
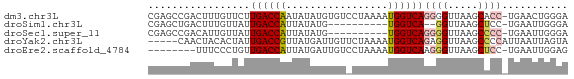
>dm3.chr3L 22751922 69 - 24543557 CGAGCCGACUUUGUUCUUGACCAAUAUAUGUGUCCUAAAAUGGUCAGGGGUUAAGCACC-UGAACUGGGA ......(.(((.(..((((((((.................))))))))..).)))).((-(.....))). ( -16.13, z-score = -0.08, R) >droSim1.chr3L 22040030 57 - 22553184 CGAGCUGACUUUGUUAUUGACCAUUAUAUG----------UGGUCA--GGUUAAGCUCC-UGAAUUGGGA .(((((((((.......(((((((.....)----------))))))--)))).))))).-.......... ( -17.01, z-score = -2.50, R) >droSec1.super_11 2672551 59 - 2888827 CGAGCCGACAUUGUUAUUGACCAUUAUAUG----------UGGUCAGGGGUUAAGCCCC-UGAAUUGGGA ....((((..........((((((.....)----------)))))(((((.....))))-)...)))).. ( -20.60, z-score = -2.52, R) >droYak2.chr3L 23243090 65 - 24197627 -----CAACUACACUAUUGACCGUUAUGAUUGUUCUAAAAUGGUCAGAGGUUAAGCCCCAUUAAUUAGUA -----.......(((((((((((((..((....))...))))))))).((.....))........)))). ( -13.20, z-score = -1.54, R) >droEre2.scaffold_4784 22403994 61 - 25762168 --------UUUCCCUGUUGACCAUUAUGAUUGUCCUAAAAUGGUCAAGGGUUAAGCUCC-UGAAUUGGAG --------...((((..((((((((.((.......)).)))))))))))).....((((-......)))) ( -18.20, z-score = -2.66, R) >consensus CGAGCCGACUUUGUUAUUGACCAUUAUAUGUGU_CUAAAAUGGUCAGGGGUUAAGCCCC_UGAAUUGGGA ........(((..((.(((((((.................))))))).))..)))............... ( -6.61 = -6.53 + -0.08)

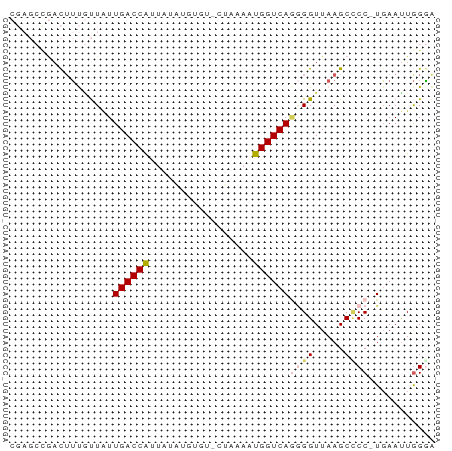
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:45:42 2011