| Sequence ID | dm3.chr3L |
|---|---|
| Location | 22,685,769 – 22,685,838 |
| Length | 69 |
| Max. P | 0.909485 |

| Location | 22,685,769 – 22,685,838 |
|---|---|
| Length | 69 |
| Sequences | 4 |
| Columns | 70 |
| Reading direction | reverse |
| Mean pairwise identity | 63.78 |
| Shannon entropy | 0.55469 |
| G+C content | 0.30343 |
| Mean single sequence MFE | -5.60 |
| Consensus MFE | -3.50 |
| Energy contribution | -2.88 |
| Covariance contribution | -0.62 |
| Combinations/Pair | 1.50 |
| Mean z-score | -1.13 |
| Structure conservation index | 0.62 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.21 |
| SVM RNA-class probability | 0.909485 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
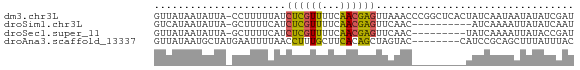
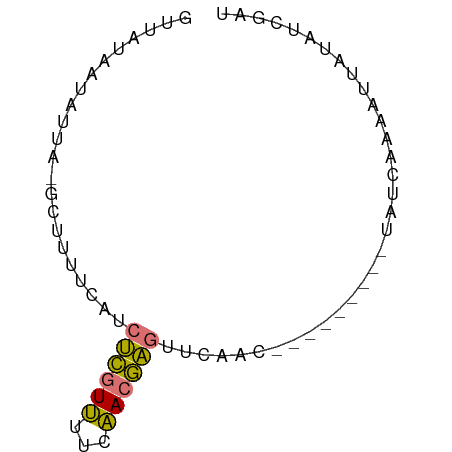
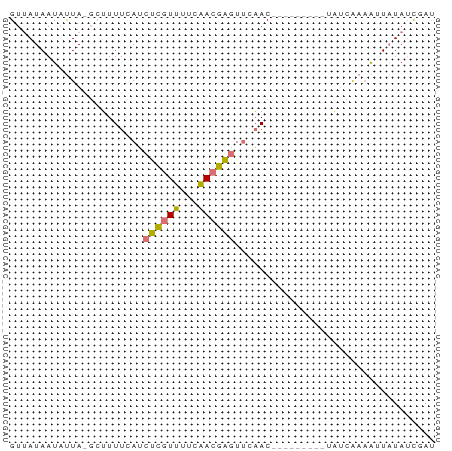
>dm3.chr3L 22685769 69 - 24543557 GUUAUAAUAUUA-CCUUUUUAUCUCGUUUUCAACGAGUUAAACCCGGCUCACUAUCAAUAAUAUAUCGAU ............-((..((((.(((((.....))))).))))...))....................... ( -7.10, z-score = -1.00, R) >droSim1.chr3L 577515 59 + 22553184 GUCAUAAUAUUA-GCUUUUCAUCUCGUUUUCAACGAGUUCAAC----------AUCAAAAUUAUAUCAAU (..(((((....-.........(((((.....)))))......----------......)))))..)... ( -4.81, z-score = -2.85, R) >droSec1.super_11 2602499 60 - 2888827 GUUAUAAUAUUA-GCUUUUCAUCUCGUUUUCAACGAGUUCAAC---------UAUCAAAAUUAUACCGAU (.((((((....-.........(((((.....)))))......---------.......)))))).)... ( -6.57, z-score = -1.84, R) >droAna3.scaffold_13337 872521 62 + 23293914 GUUAUAAUGCUAUGAAUUUUAACCUUUGCUUCACAGCUAGUAC--------CAUCCGCAGCUUUAUUUAC ........(((.((((.............)))).)))......--------................... ( -3.92, z-score = 1.19, R) >consensus GUUAUAAUAUUA_GCUUUUCAUCUCGUUUUCAACGAGUUCAAC_________UAUCAAAAUUAUAUCGAU ......................((((((...))))))................................. ( -3.50 = -2.88 + -0.62)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:45:31 2011