
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 22,651,913 – 22,651,968 |
| Length | 55 |
| Max. P | 0.978566 |

| Location | 22,651,913 – 22,651,968 |
|---|---|
| Length | 55 |
| Sequences | 7 |
| Columns | 57 |
| Reading direction | forward |
| Mean pairwise identity | 66.41 |
| Shannon entropy | 0.68270 |
| G+C content | 0.37306 |
| Mean single sequence MFE | -10.23 |
| Consensus MFE | -6.37 |
| Energy contribution | -5.99 |
| Covariance contribution | -0.38 |
| Combinations/Pair | 1.90 |
| Mean z-score | -0.98 |
| Structure conservation index | 0.62 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 2.00 |
| SVM RNA-class probability | 0.978566 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
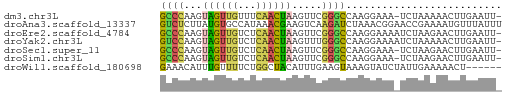
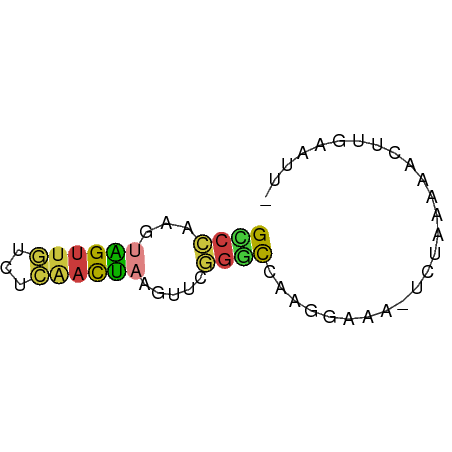
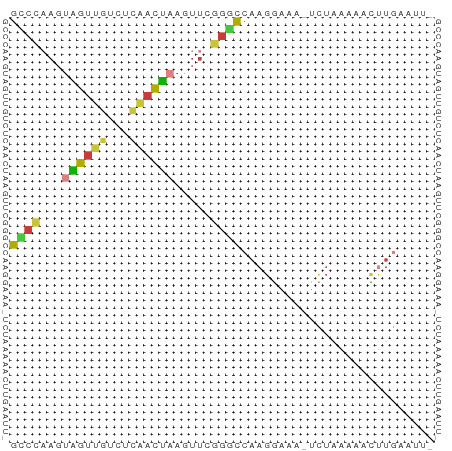
>dm3.chr3L 22651913 55 + 24543557 GCCCAAGUAGUUGUUUCAACUAAGUUCGGGCCAAGGAAA-UCUAAAAACUUGAAUU- ((((...((((((...)))))).....))))((((....-........))))....- ( -13.00, z-score = -2.07, R) >droAna3.scaffold_13337 22366536 57 + 23293914 GUCUCUUAUGUGCCAUAAACGAAGUCAAGAUCUAAACGGAACCGAAAAUGUUUAUUU ..............(((((((...((..(.(((....))).).))...))))))).. ( -3.60, z-score = 1.27, R) >droEre2.scaffold_4784 22301948 56 + 25762168 GCCCAAGUAGUUGUCUCAACUAAGUUCGGGCCAAGGAAAAUCUAAGAACUUGAAUU- ((((...((((((...)))))).....))))((((.............))))....- ( -12.92, z-score = -1.62, R) >droYak2.chr3L 23107715 56 + 24197627 GUCCAAGUAGUUGUCUCAACUAAGUUUGGGCCAAGGAAAAUCUAAAAACUUGAAUU- ((((((.((((((...))))))...))))))((((.............))))....- ( -13.12, z-score = -2.32, R) >droSec1.super_11 2568186 55 + 2888827 GCCCAAGUAGUUGUCUCAACUAAGUUCGGGCCAAGGAAA-UCUAAGAACUUGAAUU- ((((...((((((...)))))).....))))((((....-........))))....- ( -13.00, z-score = -1.62, R) >droSim1.chr3L 21937148 55 + 22553184 GCCCAAGUAGUUGUCUCAACUAAGUUCGGGCCAAGGAAA-UCUAAGAACUUGAAUU- ((((...((((((...)))))).....))))((((....-........))))....- ( -13.00, z-score = -1.62, R) >droWil1.scaffold_180698 4343972 51 - 11422946 GAAACAUUUGUUUUCUGGCUACAUUUGAAGUAAAGUAUCUAUUGAAAAACU------ ..........((((((((.(((.((......)).))).)))..)))))...------ ( -3.00, z-score = 1.12, R) >consensus GCCCAAGUAGUUGUCUCAACUAAGUUCGGGCCAAGGAAA_UCUAAAAACUUGAAUU_ ((((...((((((...)))))).....)))).......................... ( -6.37 = -5.99 + -0.38)

| Location | 22,651,913 – 22,651,968 |
|---|---|
| Length | 55 |
| Sequences | 7 |
| Columns | 57 |
| Reading direction | reverse |
| Mean pairwise identity | 66.41 |
| Shannon entropy | 0.68270 |
| G+C content | 0.37306 |
| Mean single sequence MFE | -9.43 |
| Consensus MFE | -5.02 |
| Energy contribution | -5.89 |
| Covariance contribution | 0.86 |
| Combinations/Pair | 1.58 |
| Mean z-score | -0.46 |
| Structure conservation index | 0.53 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.37 |
| SVM RNA-class probability | 0.661999 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
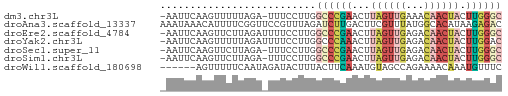
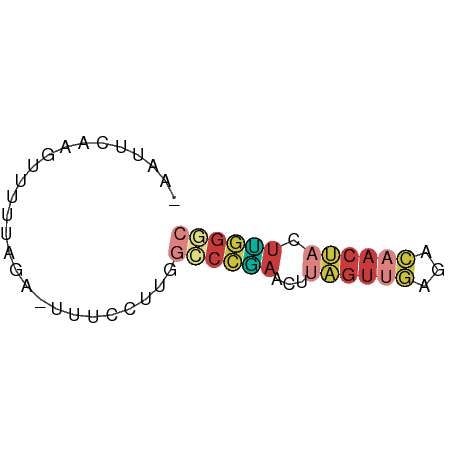
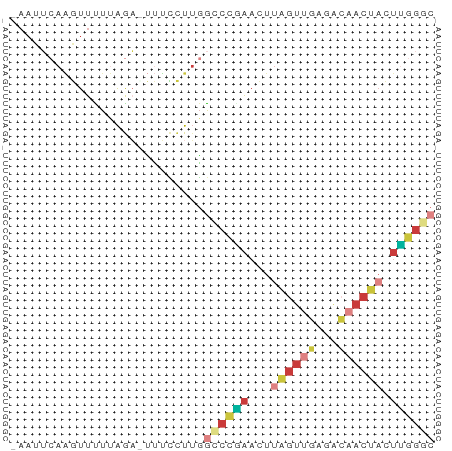
>dm3.chr3L 22651913 55 - 24543557 -AAUUCAAGUUUUUAGA-UUUCCUUGGCCCGAACUUAGUUGAAACAACUACUUGGGC -....((((........-....))))((((((...((((((...)))))).)))))) ( -12.30, z-score = -1.52, R) >droAna3.scaffold_13337 22366536 57 - 23293914 AAAUAAACAUUUUCGGUUCCGUUUAGAUCUUGACUUCGUUUAUGGCACAUAAGAGAC ..........((((.((.((((...((........))....)))).))....)))). ( -6.90, z-score = 0.28, R) >droEre2.scaffold_4784 22301948 56 - 25762168 -AAUUCAAGUUCUUAGAUUUUCCUUGGCCCGAACUUAGUUGAGACAACUACUUGGGC -....((((.............))))((((((...((((((...)))))).)))))) ( -12.22, z-score = -0.99, R) >droYak2.chr3L 23107715 56 - 24197627 -AAUUCAAGUUUUUAGAUUUUCCUUGGCCCAAACUUAGUUGAGACAACUACUUGGAC -...(((((.............))))).((((...((((((...)))))).)))).. ( -7.52, z-score = 0.19, R) >droSec1.super_11 2568186 55 - 2888827 -AAUUCAAGUUCUUAGA-UUUCCUUGGCCCGAACUUAGUUGAGACAACUACUUGGGC -....((((........-....))))((((((...((((((...)))))).)))))) ( -12.30, z-score = -1.04, R) >droSim1.chr3L 21937148 55 - 22553184 -AAUUCAAGUUCUUAGA-UUUCCUUGGCCCGAACUUAGUUGAGACAACUACUUGGGC -....((((........-....))))((((((...((((((...)))))).)))))) ( -12.30, z-score = -1.04, R) >droWil1.scaffold_180698 4343972 51 + 11422946 ------AGUUUUUCAAUAGAUACUUUACUUCAAAUGUAGCCAGAAAACAAAUGUUUC ------.((((((.......(((............)))....))))))......... ( -2.50, z-score = 0.94, R) >consensus _AAUUCAAGUUUUUAGA_UUUCCUUGGCCCGAACUUAGUUGAGACAACUACUUGGGC ..........................((((((...((((((...)))))).)))))) ( -5.02 = -5.89 + 0.86)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:45:26 2011