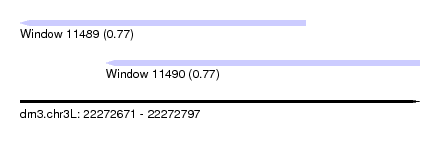

| Sequence ID | dm3.chr3L |
|---|---|
| Location | 22,272,671 – 22,272,797 |
| Length | 126 |
| Max. P | 0.768519 |

| Location | 22,272,671 – 22,272,761 |
|---|---|
| Length | 90 |
| Sequences | 12 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 62.40 |
| Shannon entropy | 0.78052 |
| G+C content | 0.54263 |
| Mean single sequence MFE | -30.86 |
| Consensus MFE | -13.85 |
| Energy contribution | -13.63 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.31 |
| Mean z-score | -0.63 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.63 |
| SVM RNA-class probability | 0.767577 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 22272671 90 - 24543557 UCAGUGGCCACUUGGCCCGCGACUUGGUCUGCUUCUCGAUGGAGCUGACUAAGGGUGGA----AC-GCAUUAGCCAUUGGCAACUCAACUUGGAU-------------- ..(((.((((..((((((((..(((((((.(((((.....))))).))))))).)))).----..-......)))).)))).)))..........-------------- ( -34.50, z-score = -2.37, R) >droGri2.scaffold_15110 14649673 109 - 24565398 UCAGCGGCCACUUGGCCCGCGUCUUCGUUUGCUUCUCAAUCGAGCUGACUAAGGAUGGGAAGGAACAAAGUGACAGAUGAUCCAACCACCAGCAAGAGCAGAAAAUACG ...((((((....)).)))).(((.(.((((((.(((....)))........((.(((...(((.((..........)).)))..))))))))))).).)))....... ( -26.50, z-score = 0.38, R) >droMoj3.scaffold_6680 2189095 97 + 24764193 UCAGCGGCCACUUGGCCCUCGUCUUCGUCUGCUUCUCGAUUGAGCUGACUGCAAGGGAAA-CAAGAGGAAGACUGAGUGGACUCUAGACUCUUUGGGC----------- .....((((....))))(((((((((.(((..(((((......((.....))..))))).-..))).)))))).((((.........))))...))).----------- ( -28.80, z-score = 0.85, R) >droVir3.scaffold_13049 7695395 99 + 25233164 UCAGCGGCCACUUGGCACGCGUCUUCGUUUGCUUCUCAAUCGAGCUGACUGUGGGCGAGG--GGAGCGAUUAGUGGGGAAUGGCAAGAAUAGCAUUUGUGC-------- ...((((((....))).))).((((((.(((((((((..(((.((.....))...)))))--)))))))....))))))...(((.((((...)))).)))-------- ( -29.20, z-score = 0.74, R) >droWil1.scaffold_180698 841297 107 - 11422946 UAAUUGGCCAUUUGGCCCGAGUCUUGGUCUGUUUCUCAAUCGAACUGACUGCAAAAGAGA-AAAGAAAAGGAGAGAUGUAAUGCUCAGAGAAAGUGGCAACUAAAUUC- ......(((((((...((...((((..(((.(((..((.((.....)).))..))).)))-.))))...)).(((........))).....)))))))..........- ( -22.60, z-score = 0.12, R) >droPer1.super_29 316758 87 - 1099123 UCAGCGGCCACUUGGACCGCGUCUUGGUCUGCUUCUCGAUGGAGCUGACUGGGGGAGGGA--GAGCAGAGGUUUAGCUGAC-GGCUGGAU------------------- ...((((((....)).)))).(((..(((.(((((.....))))).)))..)))......--........(((((((....-.)))))))------------------- ( -32.40, z-score = -0.28, R) >dp4.chrXR_group8 4084282 88 - 9212921 UCAGCGGCCACUUGGACCGCGUCUUGGUCUGCUUCUCGAUGGAGCUGACUGGGGGAGGGA--GAGCAGAGGUUUAGCUGACUGGCUGGAU------------------- ...((((((....)).)))).(((..(((.(((((.....))))).)))..)))......--........((((((((....))))))))------------------- ( -33.40, z-score = -0.47, R) >droAna3.scaffold_13337 22009992 89 - 23293914 UCAACGGCCACUUGGCCCUCGUCUUGGUUUGCUUUUCGAUUGAACUGACUGCAACCGAGA-AAGUUAGUGGGCCGCGAUGCUAUUUAUGC------------------- .....(((....((((((.(.((((((((.((...(((.......)))..))))))))))-......).))))))....)))........------------------- ( -26.20, z-score = -1.18, R) >droEre2.scaffold_4784 21943033 90 - 25762168 UGAGUGGCCACUUGGCCCGCGACUUGGUCUGCUUCUCGAUGGAGCUGACUGUAAGUGGA----AC-GCAUUAGCCAGUGGCUACUAAUGUUGGGU-------------- ..((((((((((.(((((((....(((((.(((((.....))))).)))))...)))).----..-......)))))))))))))..........-------------- ( -37.60, z-score = -2.96, R) >droYak2.chr3L 22667826 93 - 24197627 UAAGCGGCCACUUGGCCCGCGACUUGGUCUGCUUCUCGAUGGAGCUGACUGUGAGUGGAA--CGCAGUGGAAGCCAGUGGCAAAUAAGGUUGAGU-------------- ......((((((.(((((((.....((((.(((((.....))))).))))(((.......--))).))))..)))))))))..............-------------- ( -33.70, z-score = -1.17, R) >droSec1.super_11 2202751 92 - 2888827 UCAGUGGCCACUUGGCCCGCGACUUGGUCUGCUUUUCGAUGGAGCUGACUGUGGGUGGAA--CAC-GCAUUAGCCAUUGGCAACUCAAGUUGGGU-------------- .....((((....))))(.((((((((((.(((((.....))))).)))(((.(((((..--...-.......))))).)))...))))))).).-------------- ( -31.60, z-score = -0.42, R) >droSim1.chr3L 21611087 92 - 22553184 UCAGUGGCCACUUGGCCCGCGACUUGGUCUGCUUCUCGAUGGAGCUGACUGUGGGUGGAA--CAC-GCAUUAGCCAUUGGCAACUCAAGUUGGGU-------------- .....((((....))))(.((((((((((.(((((.....))))).)))(((.(((((..--...-.......))))).)))...))))))).).-------------- ( -33.80, z-score = -0.85, R) >consensus UCAGCGGCCACUUGGCCCGCGUCUUGGUCUGCUUCUCGAUGGAGCUGACUGUGGGUGGAA__GAC_GGAGUAGCCAGUGGCAACUCAAGUUGGGU______________ ...((((((....)).))))......(((.((((.......)))).)))............................................................ (-13.85 = -13.63 + -0.22)


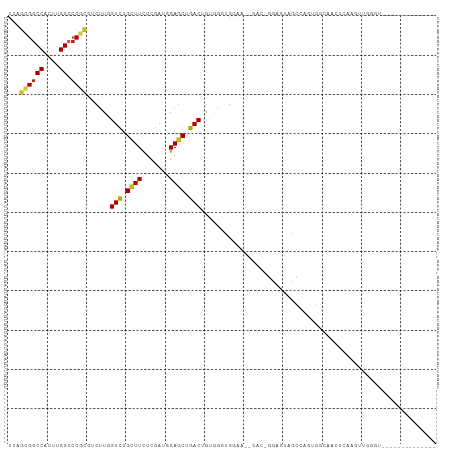
| Location | 22,272,698 – 22,272,797 |
|---|---|
| Length | 99 |
| Sequences | 11 |
| Columns | 116 |
| Reading direction | reverse |
| Mean pairwise identity | 71.60 |
| Shannon entropy | 0.56304 |
| G+C content | 0.54620 |
| Mean single sequence MFE | -35.83 |
| Consensus MFE | -17.56 |
| Energy contribution | -17.35 |
| Covariance contribution | -0.21 |
| Combinations/Pair | 1.26 |
| Mean z-score | -1.30 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.63 |
| SVM RNA-class probability | 0.768519 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
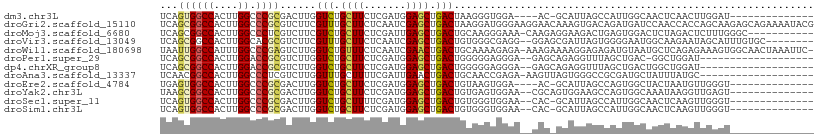
>dm3.chr3L 22272698 99 - 24543557 UGUUAGUGUUGGUGUUGGUCGAUCCAUUUGGCUCGAUCAGUGGCCACUUGGCCCGCGACUUGGUCUGCUUCUCGAUGGAGCUGACUAAGGGUGGAACGC----------------- .....(((((....((((((((.((....)).)))))))).((((....))))(((..(((((((.(((((.....))))).))))))).))).)))))----------------- ( -41.00, z-score = -2.84, R) >droGri2.scaffold_15110 14649702 107 - 24565398 ---------UGUUCGUCGUCGAGCCAUUCGGUUCGAUCAGCGGCCACUUGGCCCGCGUCUUCGUUUGCUUCUCAAUCGAGCUGACUAAGGAUGGGAAGGAACAAAGUGACAGAUGA ---------...(((((((((((((....))))))))..((((((....)).))))(((...((((.(((((((.((...........)).))))))))))).....))).))))) ( -37.70, z-score = -1.11, R) >droMoj3.scaffold_6680 2189117 99 + 24764193 ------------UGGUCGUGGAGCCAUUAGGCUCGAUCAGCGGCCACUUGGCCCUCGUCUUCGUCUGCUUCUCGAUUGAGCUGACUGCAAGGGAAACAAGAGGAAGACUGA----- ------------(((((...(((((....))))))))))..((((....))))...((((((.(((..(((((......((.....))..)))))...))).))))))...----- ( -36.80, z-score = -1.02, R) >droWil1.scaffold_180698 841329 101 - 11422946 ----------GGUAUUUGUUGAGCCAUUCGGUUCAAUAAUUGGCCAUUUGGCCCGAGUCUUGGUCUGUUUCUCAAUCGAACUGACUGCAAAAGAGAAAAGAAAAGGAGAGA----- ----------.....((((((((((....))))))))))..((((....))))....((((..(((.((((((...................)))))))))..))))....----- ( -28.01, z-score = -1.17, R) >droPer1.super_29 316776 105 - 1099123 UGUUCGUGUUGGUGUUCGUGGAUCCGUUAGGUUCGAUCAGCGGCCACUUGGACCGCGUCUUGGUCUGCUUCUCGAUGGAGCUGACUGGGGGAGGGAGAGCAGAGG----------- ..(((.((((..(.(((((((.((((...(((.((.....)))))...)))))))).(((..(((.(((((.....))))).)))..)))))).)..))))))).----------- ( -37.40, z-score = -0.04, R) >dp4.chrXR_group8 4084301 105 - 9212921 UGUUCGUGUUGGUGUUCGUGGAUCCGUUAGGUUCGAUCAGCGGCCACUUGGACCGCGUCUUGGUCUGCUUCUCGAUGGAGCUGACUGGGGGAGGGAGAGCAGAGG----------- ..(((.((((..(.(((((((.((((...(((.((.....)))))...)))))))).(((..(((.(((((.....))))).)))..)))))).)..))))))).----------- ( -37.40, z-score = -0.04, R) >droAna3.scaffold_13337 22010019 95 - 23293914 UGUUCGUAUUCGUGU---UGGACCCGUUUGGCUCGAUCAACGGCCACUUGGCCCUCGUCUUGGUUUGCUUUUCGAUUGAACUGACUGCAACCGAGAAA------------------ .....((..(((.((---..((....))..)).)))...))((((....))))....((((((((.((...(((.......)))..))))))))))..------------------ ( -28.10, z-score = -1.29, R) >droEre2.scaffold_4784 21943060 99 - 25762168 UGUUCGUGUUGGUAUGUGCCGAUCCAUUUGGCUCGAUGAGUGGCCACUUGGCCCGCGACUUGGUCUGCUUCUCGAUGGAGCUGACUGUAAGUGGAACGC----------------- ....(((.(((((....)))))((((((((..(((....(.((((....))))).)))...((((.(((((.....))))).)))).))))))))))).----------------- ( -33.80, z-score = -1.10, R) >droYak2.chr3L 22667856 93 - 24197627 ------UGUUGGUAUUUGUCGAUCCAUUUGGCUCGAUAAGCGGCCACUUGGCCCGCGACUUGGUCUGCUUCUCGAUGGAGCUGACUGUGAGUGGAACGC----------------- ------.........(((((((.((....)).)))))))((((((....)))((((....(((((.(((((.....))))).)))))...))))..)))----------------- ( -34.00, z-score = -1.95, R) >droSec1.super_11 2202778 101 - 2888827 UGUUCGUGUUGGUGUUGGUCGAUCCAUUUGGCUCGAUCAGUGGCCACUUGGCCCGCGACUUGGUCUGCUUUUCGAUGGAGCUGACUGUGGGUGGAACACGC--------------- ....(((((((((..(((((((.((....)).)))))))..).))).((.((((((.....((((.(((((.....))))).)))))))))).))))))).--------------- ( -39.20, z-score = -1.82, R) >droSim1.chr3L 21611114 101 - 22553184 UGUUCGUGUUGGUGUUGGUCGAUCCGUUUGGCUCGAUCAGUGGCCACUUGGCCCGCGACUUGGUCUGCUUCUCGAUGGAGCUGACUGUGGGUGGAACACGC--------------- ....(((((((((..(((((((.((....)).)))))))..).))).((.((((((.....((((.(((((.....))))).)))))))))).))))))).--------------- ( -40.70, z-score = -1.88, R) >consensus UGUUCGUGUUGGUGUUCGUCGAUCCAUUUGGCUCGAUCAGCGGCCACUUGGCCCGCGUCUUGGUCUGCUUCUCGAUGGAGCUGACUGUGGGUGGAACAC_A_______________ ....................((.((....)).)).....((((((....)).))))....(((((.((((.......)))).)))))............................. (-17.56 = -17.35 + -0.21)

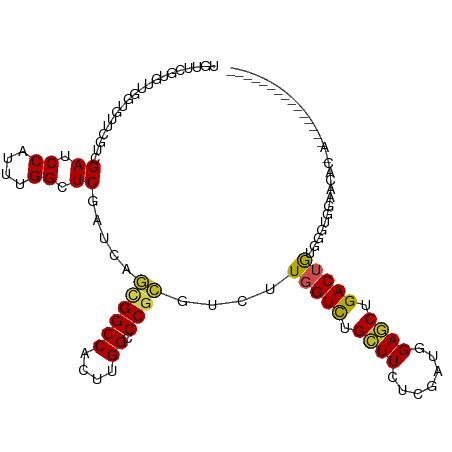

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:44:15 2011