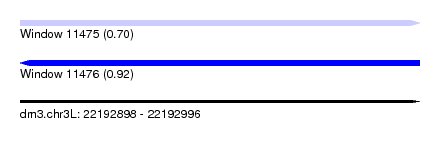

| Sequence ID | dm3.chr3L |
|---|---|
| Location | 22,192,898 – 22,192,996 |
| Length | 98 |
| Max. P | 0.920758 |

| Location | 22,192,898 – 22,192,996 |
|---|---|
| Length | 98 |
| Sequences | 10 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 84.59 |
| Shannon entropy | 0.30731 |
| G+C content | 0.29694 |
| Mean single sequence MFE | -17.69 |
| Consensus MFE | -11.67 |
| Energy contribution | -11.83 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.83 |
| Structure conservation index | 0.66 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.696949 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
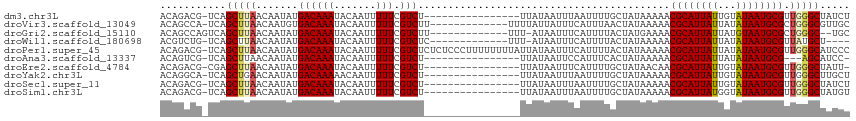
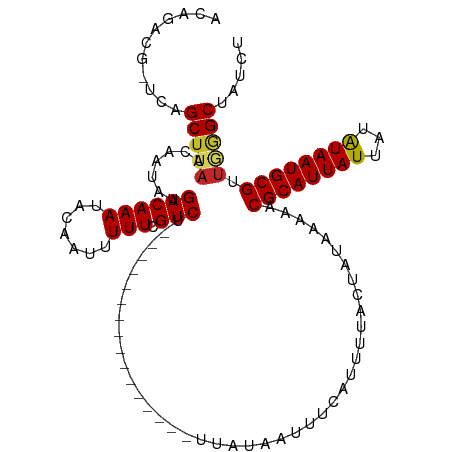
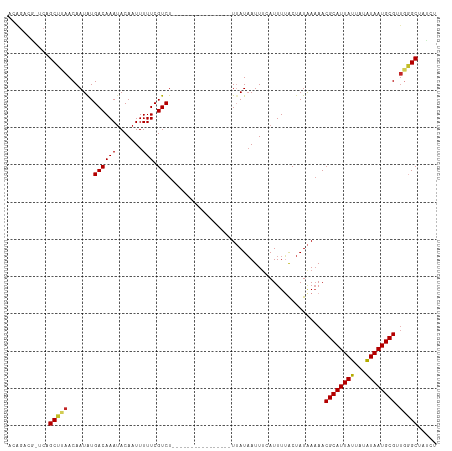
>dm3.chr3L 22192898 98 + 24543557 ACAGACG-UCAGCUUAACAAUAUGACAAAUACAAUUUUUCGUCU----------------UUAUAAUUUAAUUUUGCUAUAAAAACGCAUUAUUGUAUAAUGCGUUGGGCUAUCU .......-..((((((((......................((.(----------------(((((............)))))).))(((((((...))))))))))))))).... ( -17.60, z-score = -1.53, R) >droVir3.scaffold_13049 14755929 101 + 25233164 ACAGCCA-UCAGCUUAACAAUGUGACAAAUACAAUUUUUCGUCUU-------------UUUUAUUAUUUCAUUUAACUAUAAAAACGCAUUAUUAUAUAAUGCGCUGGGCGUUGC ...(((.-.((((..........((((((.......))).))).(-------------((((((..((......))..))))))).(((((((...))))))))))))))..... ( -17.50, z-score = -1.27, R) >droGri2.scaffold_15110 6599488 99 - 24565398 ACAGCCAGUCAGCUUAACAAUAUGACAAAUACAAUUUUUCGUCUU-------------UUU-AUAAUUUCAUUUUACUAUGAAAACGCAUUAUUAUGUAAUGCGCUGGGC--UGC .(((((...((((..........((((((.......))).)))..-------------...-....((((((......))))))..(((((((...))))))))))))))--)). ( -23.20, z-score = -3.03, R) >droWil1.scaffold_180698 109116 96 + 11422946 ACGUCUG-UCAGCUUAACAAUAUGACAAAUACAAUUUUUCGUCUC-------------UUU-AUAAUUUCAUUUUACUAUAAAAACGCAUUAUUAUAUAAUGCGUUAUGCU---- .....((-(((...........)))))..................-------------...-.....................((((((((((...)))))))))).....---- ( -13.40, z-score = -1.63, R) >droPer1.super_45 410381 114 - 618639 ACAGACG-UCAGCUUAACAAUAUGACAAAUACAAUUUUUCGUCUCUCUCCCUUUUUUUUAUUAUAAUUUCAUUUUACUAUAAAAACGCAUUAUUAUAUAAUGCGUUGGGCAUCCC ..(((((-(((...........))))(((......)))..))))....(((.....(((((..(((.......)))..)))))((((((((((...)))))))))))))...... ( -18.40, z-score = -3.06, R) >droAna3.scaffold_13337 21355172 94 + 23293914 ACAGUCG-UCAGCUUAACAAUAUGACAAAUACAAUUUUUCGUCU----------------UUAUAAUUCCAUUUCACUAUAAAAACGCAUUAUUAUAUAAUGCG---AGCAUCC- .......-...((((.........................((.(----------------(((((............)))))).))(((((((...))))))))---)))....- ( -12.10, z-score = -1.72, R) >droEre2.scaffold_4784 21864366 97 + 25762168 ACAGACG-CGAGCUUAACAAUAUGACAAAUACAAUUUUUCGUCU----------------UUAUAAUUUCAUUUUGCUAUAACAACGCAUUAUUGUAUAAUGCGUUGGGCUAUU- .......-..((((.........((((((.......))).))).----------------......................(((((((((((...)))))))))))))))...- ( -17.80, z-score = -1.47, R) >droYak2.chr3L 22577227 98 + 24197627 ACAGGCA-UCAGCUGAACAAUAUGACAAAAACAAUUUUUCGUCU----------------UUAUAAUUUAAUUUUGCUAUAAAAACGCAUUAUUGUAUAAUGCGUUGGGCUUGCU ...((((-..((((.(((.....(((.((((....)))).))).----------------..........................(((((((...)))))))))).)))))))) ( -19.80, z-score = -1.50, R) >droSec1.super_11 2123264 98 + 2888827 ACAGACG-UCAGCUUAACAAUAUGACAAAUACAAUUUUUCGUCU----------------UUAUAAUUUAAUUUUGCUAUAAAAACGCAUUAUUGUAUAAUGCGUUGGGCUAUCU .......-..((((((((......................((.(----------------(((((............)))))).))(((((((...))))))))))))))).... ( -17.60, z-score = -1.53, R) >droSim1.chr3L 21543003 98 + 22553184 ACAGACG-UCAGCUUAACAAUAUGACAAAUACAAUUUUUCGUCU----------------UUAUAAUUUAAUUUUGCUAUAAAAACGCAUUAUGGUAUAAUGCGUUGGGCUAUGU ....(((-(.((((((((......................((.(----------------(((((............)))))).))(((((((...))))))))))))))))))) ( -19.50, z-score = -1.53, R) >consensus ACAGACG_UCAGCUUAACAAUAUGACAAAUACAAUUUUUCGUCU________________UUAUAAUUUCAUUUUACUAUAAAAACGCAUUAUUAUAUAAUGCGUUGGGCUAUCU ...........(((((.......((((((.......))).)))..........................................((((((((...)))))))).)))))..... (-11.67 = -11.83 + 0.16)
| Location | 22,192,898 – 22,192,996 |
|---|---|
| Length | 98 |
| Sequences | 10 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 84.59 |
| Shannon entropy | 0.30731 |
| G+C content | 0.29694 |
| Mean single sequence MFE | -21.77 |
| Consensus MFE | -14.61 |
| Energy contribution | -14.77 |
| Covariance contribution | 0.16 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.24 |
| Structure conservation index | 0.67 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.28 |
| SVM RNA-class probability | 0.920758 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
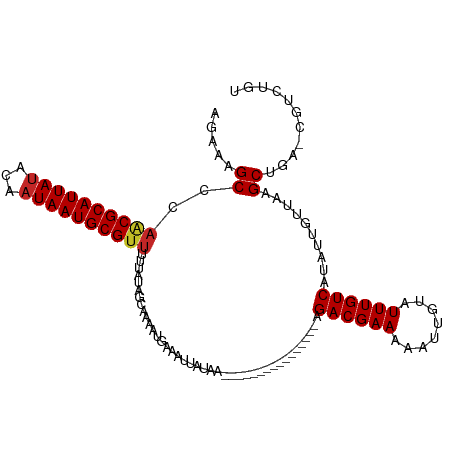
>dm3.chr3L 22192898 98 - 24543557 AGAUAGCCCAACGCAUUAUACAAUAAUGCGUUUUUAUAGCAAAAUUAAAUUAUAA----------------AGACGAAAAAUUGUAUUUGUCAUAUUGUUAAGCUGA-CGUCUGU (((((((..((((((((((...))))))))))....((((((..(((.....)))----------------.((((((........))))))...)))))).)))..-.)))).. ( -19.20, z-score = -1.65, R) >droVir3.scaffold_13049 14755929 101 - 25233164 GCAACGCCCAGCGCAUUAUAUAAUAAUGCGUUUUUAUAGUUAAAUGAAAUAAUAAAA-------------AAGACGAAAAAUUGUAUUUGUCACAUUGUUAAGCUGA-UGGCUGU .....((((((((((((((...))))))).(((((((.(((......))).))))))-------------).((((((........))))))..........)))).-.)))... ( -22.60, z-score = -1.86, R) >droGri2.scaffold_15110 6599488 99 + 24565398 GCA--GCCCAGCGCAUUACAUAAUAAUGCGUUUUCAUAGUAAAAUGAAAUUAU-AAA-------------AAGACGAAAAAUUGUAUUUGUCAUAUUGUUAAGCUGACUGGCUGU (((--(((((((((((((.....))))))..((((((......))))))....-...-------------..((((((........))))))..........))))...)))))) ( -27.10, z-score = -3.50, R) >droWil1.scaffold_180698 109116 96 - 11422946 ----AGCAUAACGCAUUAUAUAAUAAUGCGUUUUUAUAGUAAAAUGAAAUUAU-AAA-------------GAGACGAAAAAUUGUAUUUGUCAUAUUGUUAAGCUGA-CAGACGU ----(((.(((((((((((...))))))).((((((((((........)))))-)))-------------))((((((........)))))).....)))).)))..-....... ( -21.00, z-score = -2.46, R) >droPer1.super_45 410381 114 + 618639 GGGAUGCCCAACGCAUUAUAUAAUAAUGCGUUUUUAUAGUAAAAUGAAAUUAUAAUAAAAAAAAGGGAGAGAGACGAAAAAUUGUAUUUGUCAUAUUGUUAAGCUGA-CGUCUGU .((((((((((((((((((...)))))))))).(((((((........))))))).........))).....((((((........))))))...............-))))).. ( -25.10, z-score = -2.87, R) >droAna3.scaffold_13337 21355172 94 - 23293914 -GGAUGCU---CGCAUUAUAUAAUAAUGCGUUUUUAUAGUGAAAUGGAAUUAUAA----------------AGACGAAAAAUUGUAUUUGUCAUAUUGUUAAGCUGA-CGACUGU -((((((.---.(((((((...)))))))(((((((((((........)))))))----------------))))........))))))((((...........)))-)...... ( -20.40, z-score = -1.79, R) >droEre2.scaffold_4784 21864366 97 - 25762168 -AAUAGCCCAACGCAUUAUACAAUAAUGCGUUGUUAUAGCAAAAUGAAAUUAUAA----------------AGACGAAAAAUUGUAUUUGUCAUAUUGUUAAGCUCG-CGUCUGU -....((.(((((((((((...)))))))))))...((((((.((((...(((((----------------..........)))))....)))).)))))).....)-)...... ( -22.00, z-score = -2.39, R) >droYak2.chr3L 22577227 98 - 24197627 AGCAAGCCCAACGCAUUAUACAAUAAUGCGUUUUUAUAGCAAAAUUAAAUUAUAA----------------AGACGAAAAAUUGUUUUUGUCAUAUUGUUCAGCUGA-UGCCUGU .(((.((..((((((((((...))))))))))....((((..(((...((.((((----------------((((((....)))))))))).))...)))..)))).-.)).))) ( -22.20, z-score = -2.46, R) >droSec1.super_11 2123264 98 - 2888827 AGAUAGCCCAACGCAUUAUACAAUAAUGCGUUUUUAUAGCAAAAUUAAAUUAUAA----------------AGACGAAAAAUUGUAUUUGUCAUAUUGUUAAGCUGA-CGUCUGU (((((((..((((((((((...))))))))))....((((((..(((.....)))----------------.((((((........))))))...)))))).)))..-.)))).. ( -19.20, z-score = -1.65, R) >droSim1.chr3L 21543003 98 - 22553184 ACAUAGCCCAACGCAUUAUACCAUAAUGCGUUUUUAUAGCAAAAUUAAAUUAUAA----------------AGACGAAAAAUUGUAUUUGUCAUAUUGUUAAGCUGA-CGUCUGU ...((((..((((((((((...))))))))))....((((((..(((.....)))----------------.((((((........))))))...)))))).)))).-....... ( -18.90, z-score = -1.73, R) >consensus AGAAAGCCCAACGCAUUAUACAAUAAUGCGUUUUUAUAGCAAAAUGAAAUUAUAA________________AGACGAAAAAUUGUAUUUGUCAUAUUGUUAAGCUGA_CGUCUGU .....((..((((((((((...))))))))))........................................((((((........))))))..........))........... (-14.61 = -14.77 + 0.16)

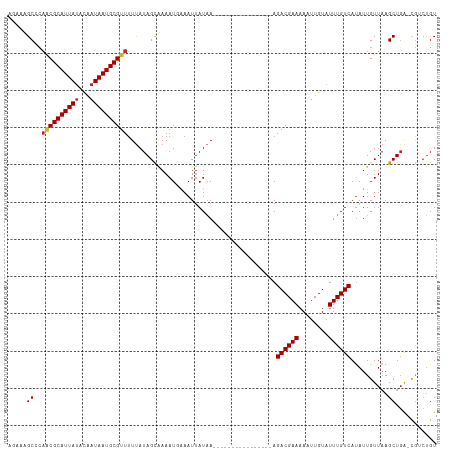
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:44:03 2011