| Sequence ID | dm3.chr3L |
|---|---|
| Location | 22,124,410 – 22,124,516 |
| Length | 106 |
| Max. P | 0.937736 |

| Location | 22,124,410 – 22,124,516 |
|---|---|
| Length | 106 |
| Sequences | 7 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 72.99 |
| Shannon entropy | 0.49422 |
| G+C content | 0.49609 |
| Mean single sequence MFE | -31.58 |
| Consensus MFE | -15.56 |
| Energy contribution | -15.34 |
| Covariance contribution | -0.22 |
| Combinations/Pair | 1.24 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.49 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.42 |
| SVM RNA-class probability | 0.937736 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
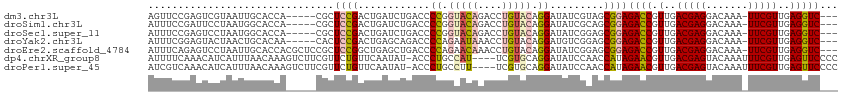
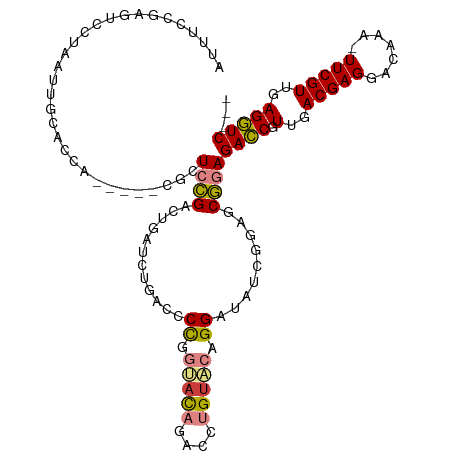
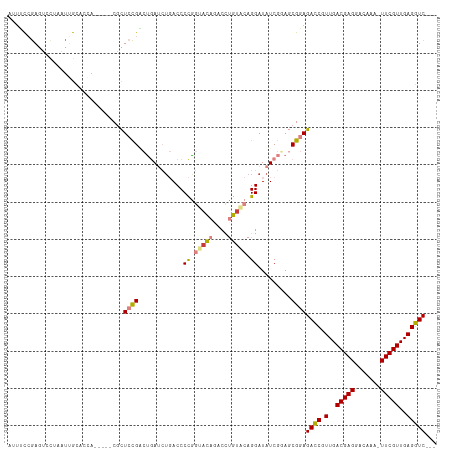
>dm3.chr3L 22124410 106 - 24543557 AGUUCCGAGUCGUAAUUGCACCA-----CGCUCCGACUGAUCUGACCCCGGUACAGACCUGUACAGGAUAUCGUAGCGGAGACCGUUGACGAGGACAAA-UUCGUUGAGGUC--- ...(((((((((.....((....-----.))..))))).........((.(((((....))))).)).........))))((((.(..(((((......-)))))..)))))--- ( -31.40, z-score = -0.30, R) >droSim1.chr3L 21471452 106 - 22553184 AUUUCCGAUUCCUAAUGGCACCA-----CGCUCCGACUGAUCUGACCCCGGUACAGACCUGUACAGGAUAUCGCAGCGGAGACCGUUGACGAGGACAAA-UUCGUUGAGGUC--- ...(((....((....)).....-----.(((.(((....((((((....)).))))(((....)))...))).))))))((((.(..(((((......-)))))..)))))--- ( -30.70, z-score = -0.65, R) >droSec1.super_11 2053770 106 - 2888827 AUUUCCGAGUCCUAAUGGCACCA-----CGCUCCGACUGAUCUGACCCCGGUACAGACCUGUACAGGAUAUCGGAGCGGAGACCGUUGACGAGGACAAA-UUCGUUGAGGUC--- ...((((.((((....)).))).-----.(((((((....((((((....)).))))(((....)))...))))))))))((((.(..(((((......-)))))..)))))--- ( -39.30, z-score = -2.47, R) >droYak2.chr3L 22468141 106 - 24197627 AUUUCGGAGUACUAACUGCACAA-----CACUCCGACUGAGCAGACCCCAGAAUAAACCUGUACAGGAUGUCGGAGCGGAGACCGUUGACGAGGACAAA-UUCGUUGAGGUC--- .....(.(((....))).)....-----(.(((((((((.((((..............)))).))....))))))).)..((((.(..(((((......-)))))..)))))--- ( -29.84, z-score = -0.98, R) >droEre2.scaffold_4784 21795346 111 - 25762168 AUUUCAGAGUCCUAAUUGCACCACGCUCCGCUCCGGCUGAGCUGACCCCAGAACAAACCUGUACAGGAUAUCGGAGCGGAGACCGUUGACGAGGACAAA-UUCGUUGAGGUC--- .................((.....))((((((((((.((..(((....)))..))..(((....)))...))))))))))((((.(..(((((......-)))))..)))))--- ( -37.30, z-score = -2.04, R) >dp4.chrXR_group8 6193747 110 - 9212921 AUUUUCAAACAUCAUUUAACAAAGUCUUCGUUCUGUUCAAUAU-ACCCUGCCAU----UCGUGCAGGAUAUCCAACCAUAGAACGUUGACGAGUACAAAUUUCGUUGAGUUCCCC .............(((((((...(((..((((((((.....((-(.((((((..----..).))))).)))......))))))))..)))(((.......))))))))))..... ( -26.30, z-score = -4.17, R) >droPer1.super_45 328197 110 + 618639 AUCGUCAAACAUCAUUUAACAAAGUCUUCGUUCUGUUCAAUAU-ACCCUGCCUU----UCGUGCAGGAUAUCCAACCAUAGAACGUUGACGAGUACAAAUUUCGUUGAGUUCCCC .............(((((((...(((..((((((((.....((-(.((((((..----..).))))).)))......))))))))..)))(((.......))))))))))..... ( -26.20, z-score = -3.64, R) >consensus AUUUCCGAGUCCUAAUUGCACCA_____CGCUCCGACUGAUCUGACCCCGGUACAGACCUGUACAGGAUAUCGGAGCGGAGACCGUUGACGAGGACAAA_UUCGUUGAGGUC___ ...............................((((............((.(((((....))))).)).........))))((((.(..(((((.......)))))..)))))... (-15.56 = -15.34 + -0.22)
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:43:51 2011