
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 22,106,167 – 22,106,257 |
| Length | 90 |
| Max. P | 0.594952 |

| Location | 22,106,167 – 22,106,257 |
|---|---|
| Length | 90 |
| Sequences | 7 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 64.65 |
| Shannon entropy | 0.65595 |
| G+C content | 0.46366 |
| Mean single sequence MFE | -12.62 |
| Consensus MFE | -5.70 |
| Energy contribution | -5.70 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -0.87 |
| Structure conservation index | 0.45 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.21 |
| SVM RNA-class probability | 0.594952 |
| Prediction | RNA |
| WARNING | Out of training range. z-scores are NOT reliable. |
Download alignment: ClustalW | MAF
>dm3.chr3L 22106167 90 - 24543557 ------------------GCUCUAUCCACCUCCAUCGCGCCUUUCCUCCCCAGCACUUCUUGCACUUCUCAUCAAUGCAAAAGACGUUUGCAUAAAGCUCUUCAAAUG-- ------------------..................((..............(((.....)))...........(((((((.....)))))))...))..........-- ( -9.90, z-score = -1.03, R) >droSec1.super_11 2035691 89 - 2888827 ------------------GCUCUGUUCACCUCCAUCGCGCCAGUCCUCCCCAGCACUUCUUGCA-UUCUCAUCAAUGCAAAAGACGUUUGCAUAAAGCUCUUCAAAUG-- ------------------(((.(((.((........((..............)).(((.(((((-((......)))))))))).....)).))).)))..........-- ( -11.64, z-score = -0.60, R) >droEre2.scaffold_4784 21777584 89 - 25762168 ------------------GUUCUAUCCACCUCCAUCGAGCCACUCCCCCCCAGCACUUCUUGCA-UUCUCAUCAAUGCAAAAGACGUUUGCAUAAAGCUCUUCAAAUG-- ------------------..................((((............(((.....))).-.........(((((((.....)))))))...))))........-- ( -13.50, z-score = -2.94, R) >droAna3.scaffold_13337 21266645 97 - 23293914 ----------GUUGCCCUCAGCUCUGCCUCCCCCGCCAGCCAUCUGCCGUGGUCUUUUAUUUCA-UUCUCAUCAAUGCAAAAGACGUUUGCAUAAAGCUCGUCAAAAG-- ----------((((....)))).........((((.(((....))).)).)).(((((.(((((-((......)))).))).((((...((.....)).)))))))))-- ( -13.60, z-score = 0.21, R) >dp4.chrXR_group8 6175031 108 - 9212921 GACCUCUCUCGCUCGCCAACGCU-CCCCCCUGCCCCCUCUUGCCGCCAGUUCGCUCUUUUUGCA-UUCUCAUCAAUGCAAAAGACGUUUGCAUAAAGCUCUUCAAAUUGA ..........(((.((.((((.(-(...................((......))...(((((((-((......))))))))))))))).))....)))............ ( -15.90, z-score = -0.52, R) >droPer1.super_45 309308 109 + 618639 GACCUCUCUCGCUCGCCAACGCUCCCCCCCUCCCCCCUCUCGCCGCCAGUUCGCUCUUUUUGCA-UUCUCAUCAAUGCAAAAGACGUUUGCAUAAAGCUCUUCAAAUUGA ..........(((.((.((((.((....................((......))...(((((((-((......))))))))))))))).))....)))............ ( -15.90, z-score = -1.39, R) >droWil1.scaffold_180955 2852273 101 + 2875958 -------AUUUUUGUCCGCUUCUCUUCGUUAUUCUUUUUUUUUAUUCUUAUUUUUUUUUUCGCA-AUUUCAUCAAUGCAAAAGACGUUUGCAUAAAGCUUUUCAAAAUG- -------..(((((...((((......(.....)..............................-.........(((((((.....))))))).))))....)))))..- ( -7.90, z-score = 0.18, R) >consensus ____________U__CC_GCUCUCUCCACCUCCCUCCUGCCACCCCCCCCCAGCUCUUCUUGCA_UUCUCAUCAAUGCAAAAGACGUUUGCAUAAAGCUCUUCAAAUG__ ..........................................................................(((((((.....)))))))................. ( -5.70 = -5.70 + 0.00)

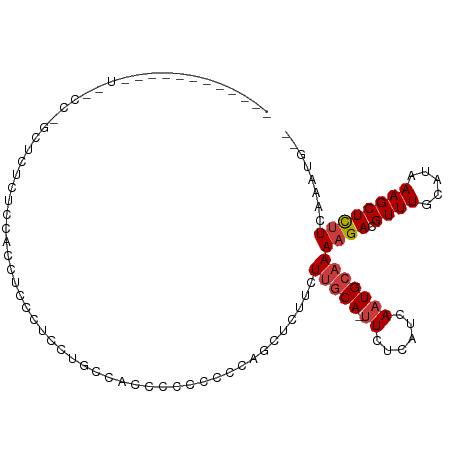

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:43:43 2011