| Sequence ID | dm3.chr3L |
|---|---|
| Location | 22,097,986 – 22,098,082 |
| Length | 96 |
| Max. P | 0.880208 |

| Location | 22,097,986 – 22,098,082 |
|---|---|
| Length | 96 |
| Sequences | 4 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 51.27 |
| Shannon entropy | 0.82382 |
| G+C content | 0.48829 |
| Mean single sequence MFE | -27.38 |
| Consensus MFE | -10.48 |
| Energy contribution | -13.10 |
| Covariance contribution | 2.62 |
| Combinations/Pair | 1.68 |
| Mean z-score | -0.80 |
| Structure conservation index | 0.38 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.42 |
| SVM RNA-class probability | 0.687824 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
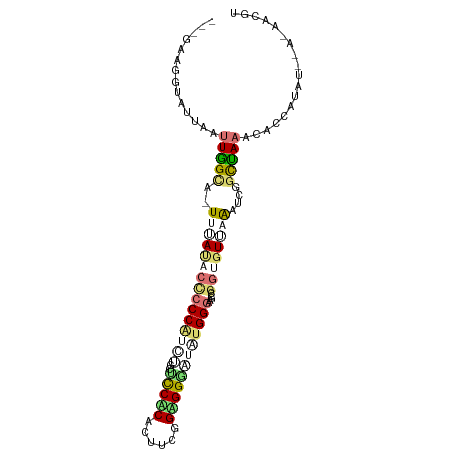
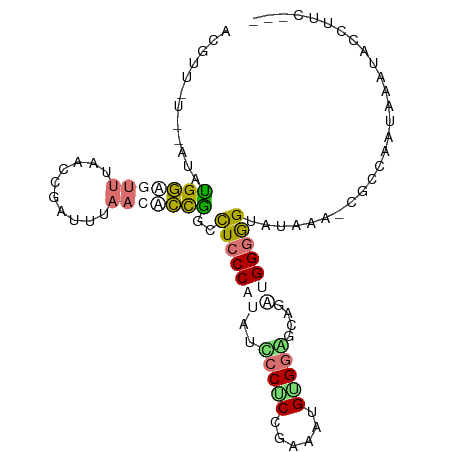
>dm3.chr3L 22097986 96 + 24543557 ---GAAGGGAUUUCUUGGCACUUUAUACCCCCGUCUGCUCCACAUUUCGGAGGGAUGUGGGAAGCUGGAGUUCGCUCAAUUAAACUCCAUAU--GAAACGA ---(..(((....)))..)..(((((((((.(((((.((((.......))))))))).)))....(((((((..........))))))))))--))).... ( -25.80, z-score = -0.27, R) >droYak2.chrU 23565886 88 + 28119190 AAAGCGGAUGCGAAUUUGCG-UUCACACUCCCUCGGGCUCCGCACAUGGGCGGACUAUGGGAGGCAGGUGUGAAAACUGCAAAGCACCU------------ ...(((((((((....))))-)))...(((((...((.(((((......)))))))..)))))))((((((............))))))------------ ( -35.30, z-score = -2.49, R) >apiMel3.GroupUn 219586076 96 - 399230636 ---GAAAGCAUUGAUUGGUGAUUUAUACCACCAUAUGCAUCACAUUUCGGUGGCUUAUGGGAAAGCACUGUUAAAUCGGCUAAAUGACACAU--AUUACGU ---....((((....(((((........))))).))))...........(((((((......)))).(((......)))........)))..--....... ( -17.70, z-score = 0.35, R) >triCas2.chrUn_298 5931 98 - 21235 ---GAACGUCUUCGGUGGCAUUGAAACCCCCCAUCAGCCCCUCACUUUGGAGGGAUUUGGGAGGCCGGCAUUAAGUCGGUCAAAACCCAUUUUUACAUCGU ---.........(((((............((((.....(((((......)))))...)))).(((((((.....)))))))..............))))). ( -30.70, z-score = -0.80, R) >consensus ___GAAGGUAUUAAUUGGCA_UUUAUACCCCCAUCUGCUCCACACUUCGGAGGGAUAUGGGAAGCCGGUGUUAAAUCGGCUAAACACCAUAU__A_AACGU ..............(((((..(((((((((((((((..(((((......)))))))))))).....))))))))....))))).................. (-10.48 = -13.10 + 2.62)


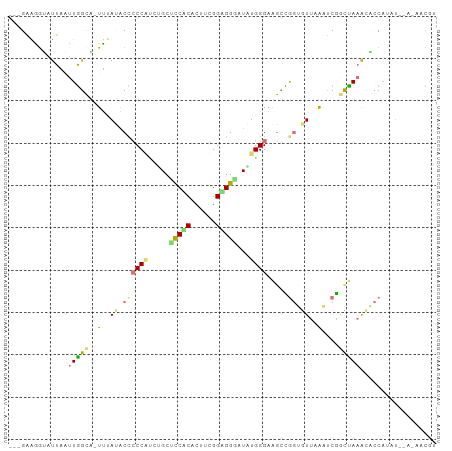
| Location | 22,097,986 – 22,098,082 |
|---|---|
| Length | 96 |
| Sequences | 4 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 51.27 |
| Shannon entropy | 0.82382 |
| G+C content | 0.48829 |
| Mean single sequence MFE | -27.48 |
| Consensus MFE | -9.61 |
| Energy contribution | -10.68 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.63 |
| Mean z-score | -1.27 |
| Structure conservation index | 0.35 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 1.04 |
| SVM RNA-class probability | 0.880208 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
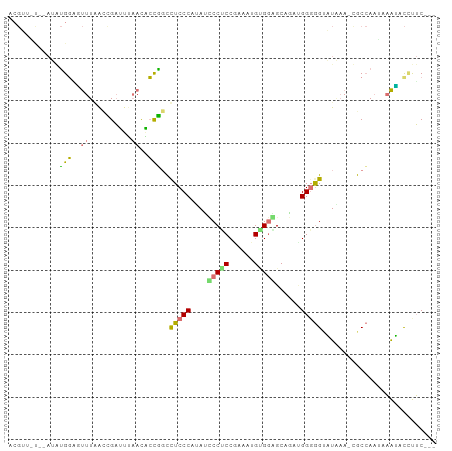
>dm3.chr3L 22097986 96 - 24543557 UCGUUUC--AUAUGGAGUUUAAUUGAGCGAACUCCAGCUUCCCACAUCCCUCCGAAAUGUGGAGCAGACGGGGGUAUAAAGUGCCAAGAAAUCCCUUC--- .(((((.--...((((((((........))))))))(((..((((((.........))))))))))))))(((((...............)))))...--- ( -25.76, z-score = -1.05, R) >droYak2.chrU 23565886 88 - 28119190 ------------AGGUGCUUUGCAGUUUUCACACCUGCCUCCCAUAGUCCGCCCAUGUGCGGAGCCCGAGGGAGUGUGAA-CGCAAAUUCGCAUCCGCUUU ------------.((((((((((.((..((((((.(.((((......(((((......)))))....)))).))))))))-))))))...)))))...... ( -30.80, z-score = -1.98, R) >apiMel3.GroupUn 219586076 96 + 399230636 ACGUAAU--AUGUGUCAUUUAGCCGAUUUAACAGUGCUUUCCCAUAAGCCACCGAAAUGUGAUGCAUAUGGUGGUAUAAAUCACCAAUCAAUGCUUUC--- ..(((((--(((((((((.....((........((((((......))).)))))....)))))))))))((((((....))))))......)))....--- ( -20.80, z-score = -1.12, R) >triCas2.chrUn_298 5931 98 + 21235 ACGAUGUAAAAAUGGGUUUUGACCGACUUAAUGCCGGCCUCCCAAAUCCCUCCAAAGUGAGGGGCUGAUGGGGGGUUUCAAUGCCACCGAAGACGUUC--- ..(((((.......(((.(((((((.(.....).)))(((((((..((((((......))))))....)))))))..)))).))).......))))).--- ( -32.54, z-score = -0.93, R) >consensus ACGUU_U__AUAUGGAGUUUAACCGAUUUAACACCGGCCUCCCAUAUCCCUCCGAAAUGUGGAGCAGAUGGGGGUAUAAA_CGCCAAUAAAUACCUUC___ ............(((((((..........)))))))..(((((((..(((((......)))))....)))))))........................... ( -9.61 = -10.68 + 1.06)

Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:43:41 2011