
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 22,076,252 – 22,076,349 |
| Length | 97 |
| Max. P | 0.807147 |

| Location | 22,076,252 – 22,076,349 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 75.53 |
| Shannon entropy | 0.39615 |
| G+C content | 0.35992 |
| Mean single sequence MFE | -24.76 |
| Consensus MFE | -12.00 |
| Energy contribution | -11.68 |
| Covariance contribution | -0.32 |
| Combinations/Pair | 1.24 |
| Mean z-score | -2.18 |
| Structure conservation index | 0.48 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.63 |
| SVM RNA-class probability | 0.767136 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
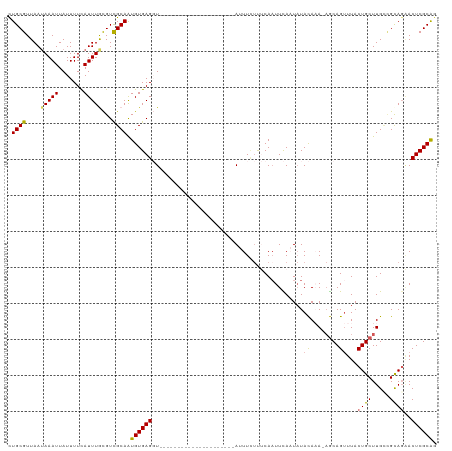
>dm3.chr3L 22076252 97 + 24543557 CUGCGUUAAUAAAUUAUUUUCAUUUGCGUCGCAAUGUGAGGU---------------------AUUUAUUUCACUUCAAUAUACACA-AGCAGUUUACUGCUUGUUGUAGAACUCGCAG (((((..(((((((...((((((((((...)))).)))))).---------------------)))))))...........((((((-(((((....))))))).)))).....))))) ( -27.50, z-score = -3.79, R) >droSim1.chr3L 21424459 97 + 22553184 CUGCGUUAAUAAAUUAUUUUUAUUUGCGUCGCAAUGUGAGGU---------------------AAUUCUUUCAAUUCACUAUACAAA-AGCAGUUUACUGCUUGCCGCAGAACUCGCGG (((((....((((.....))))((((((..((..((((.(((---------------------((((.....)))).))).)))).(-(((((....))))))))))))))...))))) ( -23.30, z-score = -1.84, R) >droSec1.super_11 2011727 96 + 2888827 CUGCGUUAAUAAAUUAUUUUUAUUUGCGUUGCAAUGUGAGGU---------------------AAUUCUUUCAAUUCACUAUACAAA-AGCAGUUUACUGCUUGC-GUAGAACUCGCGG (((((......(((((((((...((((...))))...)))))---------------------))))...........((((....(-(((((....))))))..-))))....))))) ( -22.00, z-score = -1.53, R) >droYak2.chr3L 22419910 119 + 24197627 CUGCGUUAAGAAAUUAUUUUCAUUUAUGUCGCAAUGUGAGCUUAUCUCAAAAACAAGUGGGAUAUUUUUCCCAAUUUAAUAUAUAUAUAACAGCUUACUGCUUGCCGCAUUACUCGCAC .((((....((((....))))....((((.((((.((((((((((............(((((......))))).............)))..)))))))...)))).))))....)))). ( -27.01, z-score = -2.69, R) >droEre2.scaffold_4784 21754320 106 + 25762168 UUGCGUUAAUAAAUUAUUUUCAUUUAUGUCGCAGUGUGAGCUUA----------AUGUUGAGUAUUUC-CCCAAUUUAAUCUACAAA--GCAGUUUACUGCUUGGCGCAGGACUCGCAG (((((...((((((.......))))))..)))))(((((((((.----------..(((((((.....-....))))))).....((--((((....)))))).....))).)))))). ( -24.00, z-score = -1.04, R) >consensus CUGCGUUAAUAAAUUAUUUUCAUUUGCGUCGCAAUGUGAGGU_____________________AUUUCUUUCAAUUCAAUAUACAAA_AGCAGUUUACUGCUUGCCGCAGAACUCGCAG .((((....(((((.......)))))...)))).((((((..........................................((........))...((((.....))))..)))))). (-12.00 = -11.68 + -0.32)


| Location | 22,076,252 – 22,076,349 |
|---|---|
| Length | 97 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 75.53 |
| Shannon entropy | 0.39615 |
| G+C content | 0.35992 |
| Mean single sequence MFE | -24.36 |
| Consensus MFE | -11.32 |
| Energy contribution | -11.36 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.22 |
| Mean z-score | -2.33 |
| Structure conservation index | 0.46 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.75 |
| SVM RNA-class probability | 0.807147 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
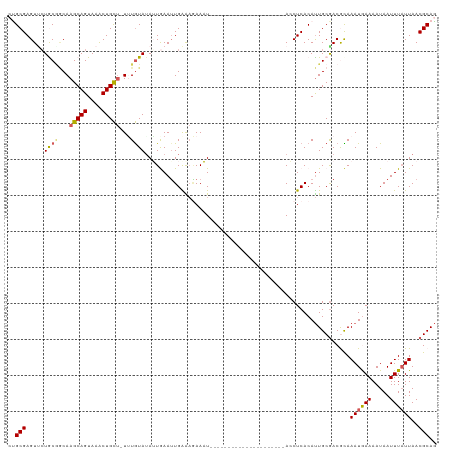
>dm3.chr3L 22076252 97 - 24543557 CUGCGAGUUCUACAACAAGCAGUAAACUGCU-UGUGUAUAUUGAAGUGAAAUAAAU---------------------ACCUCACAUUGCGACGCAAAUGAAAAUAAUUUAUUAACGCAG (((((.....((((.(((((((....)))))-))))))((.((((((((.......---------------------...)))).((((...))))..........)))).)).))))) ( -24.00, z-score = -3.10, R) >droSim1.chr3L 21424459 97 - 22553184 CCGCGAGUUCUGCGGCAAGCAGUAAACUGCU-UUUGUAUAGUGAAUUGAAAGAAUU---------------------ACCUCACAUUGCGACGCAAAUAAAAAUAAUUUAUUAACGCAG ..(((.....((((..((((((....)))))-)(((((..((((..(((.....))---------------------)..))))..)))))))))((((((.....))))))..))).. ( -23.30, z-score = -1.91, R) >droSec1.super_11 2011727 96 - 2888827 CCGCGAGUUCUAC-GCAAGCAGUAAACUGCU-UUUGUAUAGUGAAUUGAAAGAAUU---------------------ACCUCACAUUGCAACGCAAAUAAAAAUAAUUUAUUAACGCAG ..(((.....(((-(.((((((....)))))-).))))((((((((((...((...---------------------...))...((((...)))).......)))))))))).))).. ( -21.90, z-score = -2.60, R) >droYak2.chr3L 22419910 119 - 24197627 GUGCGAGUAAUGCGGCAAGCAGUAAGCUGUUAUAUAUAUAUUAAAUUGGGAAAAAUAUCCCACUUGUUUUUGAGAUAAGCUCACAUUGCGACAUAAAUGAAAAUAAUUUCUUAACGCAG .((((.((((((.(((.(((((....)))))...............(((((......)))))................)).).)))))).........((((....))))....)))). ( -26.20, z-score = -1.44, R) >droEre2.scaffold_4784 21754320 106 - 25762168 CUGCGAGUCCUGCGCCAAGCAGUAAACUGC--UUUGUAGAUUAAAUUGGG-GAAAUACUCAACAU----------UAAGCUCACACUGCGACAUAAAUGAAAAUAAUUUAUUAACGCAA .((.((((.(((((..((((((....))))--)))))))......(((((-......)))))...----------...)))).)).((((..((((((.......))))))...)))). ( -26.40, z-score = -2.59, R) >consensus CUGCGAGUUCUGCGGCAAGCAGUAAACUGCU_UUUGUAUAUUGAAUUGAAAGAAAU_____________________ACCUCACAUUGCGACGCAAAUGAAAAUAAUUUAUUAACGCAG ..(((.....((((...(((((....)))))...)))).........................................................((((((.....))))))..))).. (-11.32 = -11.36 + 0.04)


Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:43:40 2011