
| Sequence ID | dm3.chr3L |
|---|---|
| Location | 22,034,698 – 22,034,813 |
| Length | 115 |
| Max. P | 0.545026 |

| Location | 22,034,698 – 22,034,813 |
|---|---|
| Length | 115 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 77.92 |
| Shannon entropy | 0.40114 |
| G+C content | 0.44958 |
| Mean single sequence MFE | -23.53 |
| Consensus MFE | -12.17 |
| Energy contribution | -13.53 |
| Covariance contribution | 1.36 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.73 |
| Structure conservation index | 0.52 |
| Background model | dinucleotide |
| Decision model | sequence based alignment quality |
| SVM decision value | 0.10 |
| SVM RNA-class probability | 0.545026 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>dm3.chr3L 22034698 115 + 24543557 CUGCACACUUCUGC-CGACUUUCUUUUCGAGUCGCCCU-CUCCCUUGGCCUCGCCGGGCCAUUUUCGCGAUUUCCCUUCGUAAAUGUUUUUACCGGAAAUUGUUGCAAUUUAUGAUU .((((.........-((((((.......))))))....-......((((((....)))))).....(((((((((....(((((....))))).))))))))))))).......... ( -28.70, z-score = -2.40, R) >droPer1.super_45 243812 90 - 618639 UUGAACAUUUCACUUCGACUUUCUUUUCG---CGCGCU------------------------UCGCGCGCUUCGCUUUGGUGAAUGUUUUUACCGGAAAUUGUUGCAAUUUAUGAUU (((((((((((.................(---(((((.------------------------..)))))).......(((((((....))))))))))).)))).)))......... ( -22.40, z-score = -1.43, R) >droEre2.scaffold_4784 21710086 103 + 25762168 CAGCGCUUUUCACC-CGACUUUCCUUUCAAGUCGCCCU-------------CGCCCGGCCAUUUUCGCGAUUUCCCUUCGUAAAUGUUUUUACCGGAAACGGUUGCAAUUUAUGAUU ...(((........-.(((((.......)))))(((..-------------.....))).......)))........((((((((((....((((....)))).)).)))))))).. ( -25.50, z-score = -2.44, R) >droYak2.chr3L 22377204 103 + 24197627 CAGCACAUUUCUCC-GCACUUUCUUUUCAAGUCGCCCU-------------CGCCAGGCCAAUUUCGCAAUUUCCCUUCGUAAAUGUUUUUACUGGAAAUGGCUGCAAUUUAUGAUU ((((.(((((((..-((((((.......)))).(((..-------------.....))).......))...........(((((....))))).)))))))))))............ ( -19.50, z-score = -0.89, R) >droSec1.super_11 1966126 116 + 2888827 CAGCACAUUUCUGC-CGACUUUCUUCUCGAGUCGCCCUUUUCCCUUGCCAUCACCCGGCCAUUUUCGCGAUUUCCCUUCGUAAAUGUUUUUACCGGAAAUUGUUGCAAUUUAUGAUU (((((.(((((((.-((((((.......))))))............(((.......)))(((((..((((.......))))))))).......))))))))))))............ ( -22.30, z-score = -1.79, R) >droSim1.chr3L 21380208 116 + 22553184 CAGCACAUUUAUGC-CGACUUUCUUCUCGAGUCGCCCUUCUCCCUUGCCAUCGCCCGGCCAUUUUCGCGAUUUCCCUUCGUAAAUGUUUUUACCGGAAAUUGUUGCAAUUUAUGAUU ..(((.......((-((((((.......))))).............((....))..))).......(((((((((....(((((....))))).))))))))))))........... ( -22.80, z-score = -1.45, R) >consensus CAGCACAUUUCUCC_CGACUUUCUUUUCGAGUCGCCCU_____________CGCCCGGCCAUUUUCGCGAUUUCCCUUCGUAAAUGUUUUUACCGGAAAUUGUUGCAAUUUAUGAUU ..(((..........((((((.......))))))................................(((((((((....(((((....))))).))))))))))))........... (-12.17 = -13.53 + 1.36)

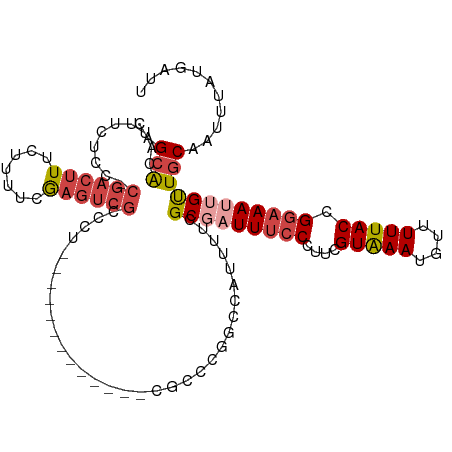
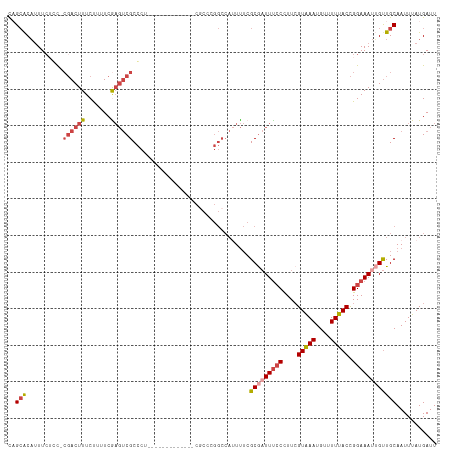
Generated by rnazCluster.pl (part of RNAz 1.0) on Tue Apr 19 23:43:34 2011